""
]
},
""metadata"": {
""needs_background"": ""light""
},
""output_type"": ""display_data""
}
],
""source"": [
""#然后通过相关分析提出自相关的变量\n"",
""cor=feature_train[choose_feature].corr()\n"",
""f , ax = plt.subplots(figsize = (9,9))\n"",
""\n"",
""\n"",
""cor""
]
},
{
""cell_type"": ""code"",
""execution_count"": 232,
""id"": ""93434e53"",
""metadata"": {},
""outputs"": [
{
""data"": {
""text/plain"": [
""(1974, 21)""
]
},
""execution_count"": 232,
""metadata"": {},
""output_type"": ""execute_result""
}
],
""source"": [
""for i in range(55):\n"",
"" for j in range(55):\n"",
"" if i !=j:\n"",
"" if cor.iloc[i,j]>0.5 and cor.index[i] in choose_feature and cor.columns[j] in choose_feature:\n"",
"" if len(choose_feature)==20:\n"",
"" break;\n"",
"" choose_feature.remove(cor.columns[j])\n"",
"" \n"",
"" \n"",
"" \n"",
"" \n"",
""choose_feature \n"",
""final_feature_train=feature_train[choose_feature]\n"",
""final_feature_train.shape""
]
},
{
""cell_type"": ""code"",
""execution_count"": 237,
""id"": ""bb2758ee"",
""metadata"": {},
""outputs"": [],
""source"": [
""final_feature_train.to_csv('final.csv') ""
]
},
{
""cell_type"": ""code"",
""execution_count"": 233,
""id"": ""101bda2e"",
""metadata"": {},
""outputs"": [],
""source"": [
""from sklearn.model_selection import cross_val_score\n"",
""from sklearn.metrics import mean_squared_error,make_scorer,mean_absolute_error\n"",
""from sklearn.linear_model import LinearRegression\n"",
""from sklearn.tree import DecisionTreeRegressor\n"",
""from sklearn.ensemble import RandomForestRegressor\n"",
""from sklearn.ensemble import GradientBoostingRegressor\n"",
""from sklearn.neural_network import MLPRegressor\n"",
""from xgboost.sklearn import XGBRegressor \n"",
""from lightgbm.sklearn import LGBMRegressor""
]
},
{
""cell_type"": ""code"",
""execution_count"": 234,
""id"": ""07923e6e"",
""metadata"": {},
""outputs"": [
{
""name"": ""stdout"",
""output_type"": ""stream"",
""text"": [
""LinearRegression is finished\n"",
""DecisionTreeRegressor is finished\n""
]
},
{
""name"": ""stderr"",
""output_type"": ""stream"",
""text"": [
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\model_selection\\_validation.py:598: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples,), for example using ravel().\n"",
"" estimator.fit(X_train, y_train, **fit_params)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\model_selection\\_validation.py:598: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples,), for example using ravel().\n"",
"" estimator.fit(X_train, y_train, **fit_params)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\model_selection\\_validation.py:598: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples,), for example using ravel().\n"",
"" estimator.fit(X_train, y_train, **fit_params)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\model_selection\\_validation.py:598: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples,), for example using ravel().\n"",
"" estimator.fit(X_train, y_train, **fit_params)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\model_selection\\_validation.py:598: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples,), for example using ravel().\n"",
"" estimator.fit(X_train, y_train, **fit_params)\n""
]
},
{
""name"": ""stdout"",
""output_type"": ""stream"",
""text"": [
""RandomForestRegressor is finished\n""
]
},
{
""name"": ""stderr"",
""output_type"": ""stream"",
""text"": [
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\utils\\validation.py:63: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples, ), for example using ravel().\n"",
"" return f(*args, **kwargs)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\utils\\validation.py:63: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples, ), for example using ravel().\n"",
"" return f(*args, **kwargs)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\utils\\validation.py:63: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples, ), for example using ravel().\n"",
"" return f(*args, **kwargs)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\utils\\validation.py:63: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples, ), for example using ravel().\n"",
"" return f(*args, **kwargs)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\utils\\validation.py:63: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples, ), for example using ravel().\n"",
"" return f(*args, **kwargs)\n""
]
},
{
""name"": ""stdout"",
""output_type"": ""stream"",
""text"": [
""GradientBoostingRegressor is finished\n""
]
},
{
""name"": ""stderr"",
""output_type"": ""stream"",
""text"": [
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\utils\\validation.py:63: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples, ), for example using ravel().\n"",
"" return f(*args, **kwargs)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\neural_network\\_multilayer_perceptron.py:500: ConvergenceWarning: lbfgs failed to converge (status=1):\n"",
""STOP: TOTAL NO. of ITERATIONS REACHED LIMIT.\n"",
""\n"",
""Increase the number of iterations (max_iter) or scale the data as shown in:\n"",
"" https://scikit-learn.org/stable/modules/preprocessing.html\n"",
"" self.n_iter_ = _check_optimize_result(\""lbfgs\"", opt_res, self.max_iter)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\utils\\validation.py:63: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples, ), for example using ravel().\n"",
"" return f(*args, **kwargs)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\neural_network\\_multilayer_perceptron.py:500: ConvergenceWarning: lbfgs failed to converge (status=1):\n"",
""STOP: TOTAL NO. of ITERATIONS REACHED LIMIT.\n"",
""\n"",
""Increase the number of iterations (max_iter) or scale the data as shown in:\n"",
"" https://scikit-learn.org/stable/modules/preprocessing.html\n"",
"" self.n_iter_ = _check_optimize_result(\""lbfgs\"", opt_res, self.max_iter)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\utils\\validation.py:63: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples, ), for example using ravel().\n"",
"" return f(*args, **kwargs)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\neural_network\\_multilayer_perceptron.py:500: ConvergenceWarning: lbfgs failed to converge (status=1):\n"",
""STOP: TOTAL NO. of ITERATIONS REACHED LIMIT.\n"",
""\n"",
""Increase the number of iterations (max_iter) or scale the data as shown in:\n"",
"" https://scikit-learn.org/stable/modules/preprocessing.html\n"",
"" self.n_iter_ = _check_optimize_result(\""lbfgs\"", opt_res, self.max_iter)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\utils\\validation.py:63: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples, ), for example using ravel().\n"",
"" return f(*args, **kwargs)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\neural_network\\_multilayer_perceptron.py:500: ConvergenceWarning: lbfgs failed to converge (status=1):\n"",
""STOP: TOTAL NO. of ITERATIONS REACHED LIMIT.\n"",
""\n"",
""Increase the number of iterations (max_iter) or scale the data as shown in:\n"",
"" https://scikit-learn.org/stable/modules/preprocessing.html\n"",
"" self.n_iter_ = _check_optimize_result(\""lbfgs\"", opt_res, self.max_iter)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\utils\\validation.py:63: DataConversionWarning: A column-vector y was passed when a 1d array was expected. Please change the shape of y to (n_samples, ), for example using ravel().\n"",
"" return f(*args, **kwargs)\n"",
""d:\\download\\minconda\\envs\\d2l-zh\\lib\\site-packages\\sklearn\\neural_network\\_multilayer_perceptron.py:500: ConvergenceWarning: lbfgs failed to converge (status=1):\n"",
""STOP: TOTAL NO. of ITERATIONS REACHED LIMIT.\n"",
""\n"",
""Increase the number of iterations (max_iter) or scale the data as shown in:\n"",
"" https://scikit-learn.org/stable/modules/preprocessing.html\n"",
"" self.n_iter_ = _check_optimize_result(\""lbfgs\"", opt_res, self.max_iter)\n""
]
},
{
""name"": ""stdout"",
""output_type"": ""stream"",
""text"": [
""MLPRegressor is finished\n"",
""[19:30:56] WARNING: ..\\src\\learner.cc:541: \n"",
""Parameters: { n_estimator } might not be used.\n"",
""\n"",
"" This may not be accurate due to some parameters are only used in language bindings but\n"",
"" passed down to XGBoost core. Or some parameters are not used but slip through this\n"",
"" verification. Please open an issue if you find above cases.\n"",
""\n"",
""\n"",
""[19:30:56] WARNING: ..\\src\\learner.cc:541: \n"",
""Parameters: { n_estimator } might not be used.\n"",
""\n"",
"" This may not be accurate due to some parameters are only used in language bindings but\n"",
"" passed down to XGBoost core. Or some parameters are not used but slip through this\n"",
"" verification. Please open an issue if you find above cases.\n"",
""\n"",
""\n"",
""[19:30:57] WARNING: ..\\src\\learner.cc:541: \n"",
""Parameters: { n_estimator } might not be used.\n"",
""\n"",
"" This may not be accurate due to some parameters are only used in language bindings but\n"",
"" passed down to XGBoost core. Or some parameters are not used but slip through this\n"",
"" verification. Please open an issue if you find above cases.\n"",
""\n"",
""\n"",
""[19:30:57] WARNING: ..\\src\\learner.cc:541: \n"",
""Parameters: { n_estimator } might not be used.\n"",
""\n"",
"" This may not be accurate due to some parameters are only used in language bindings but\n"",
"" passed down to XGBoost core. Or some parameters are not used but slip through this\n"",
"" verification. Please open an issue if you find above cases.\n"",
""\n"",
""\n"",
""[19:30:57] WARNING: ..\\src\\learner.cc:541: \n"",
""Parameters: { n_estimator } might not be used.\n"",
""\n"",
"" This may not be accurate due to some parameters are only used in language bindings but\n"",
"" passed down to XGBoost core. Or some parameters are not used but slip through this\n"",
"" verification. Please open an issue if you find above cases.\n"",
""\n"",
""\n"",
""XGBRegressor is finished\n"",
""[LightGBM] [Warning] Unknown parameter: n_estimator\n"",
""[LightGBM] [Warning] Unknown parameter: n_estimator\n"",
""[LightGBM] [Warning] Unknown parameter: n_estimator\n"",
""[LightGBM] [Warning] Unknown parameter: n_estimator\n"",
""[LightGBM] [Warning] Unknown parameter: n_estimator\n"",
""LGBMRegressor is finished\n""
]
},
{
""data"": {
""text/html"": [
""
\n"",
""\n"",
""
\n"",
"" \n"",
""
\n"",
""
\n"",
""
LinearRegression
\n"",
""
DecisionTreeRegressor
\n"",
""
RandomForestRegressor
\n"",
""
GradientBoostingRegressor
\n"",
""
MLPRegressor
\n"",
""
XGBRegressor
\n"",
""
LGBMRegressor
\n"",
""
\n"",
"" \n"",
"" \n"",
""
\n"",
""
cv1
\n"",
""
0.847049
\n"",
""
1.917903
\n"",
""
0.718402
\n"",
""
0.706266
\n"",
""
0.670726
\n"",
""
0.983154
\n"",
""
0.887868
\n"",
""
\n"",
""
\n"",
""
cv2
\n"",
""
0.719566
\n"",
""
2.064269
\n"",
""
0.627625
\n"",
""
0.632428
\n"",
""
0.837934
\n"",
""
0.586317
\n"",
""
0.649590
\n"",
""
\n"",
""
\n"",
""
cv3
\n"",
""
0.899025
\n"",
""
1.292795
\n"",
""
0.781737
\n"",
""
0.779887
\n"",
""
0.874925
\n"",
""
0.999408
\n"",
""
0.879154
\n"",
""
\n"",
""
\n"",
""
cv4
\n"",
""
2.093724
\n"",
""
2.352413
\n"",
""
1.116827
\n"",
""
1.076843
\n"",
""
1.366192
\n"",
""
1.070108
\n"",
""
1.108118
\n"",
""
\n"",
""
\n"",
""
cv5
\n"",
""
1.389543
\n"",
""
2.381274
\n"",
""
1.491859
\n"",
""
1.508647
\n"",
""
1.533531
\n"",
""
1.566166
\n"",
""
1.462994
\n"",
""
\n"",
"" \n"",
""
\n"",
""
""
],
""text/plain"": [
"" LinearRegression DecisionTreeRegressor RandomForestRegressor \\\n"",
""cv1 0.847049 1.917903 0.718402 \n"",
""cv2 0.719566 2.064269 0.627625 \n"",
""cv3 0.899025 1.292795 0.781737 \n"",
""cv4 2.093724 2.352413 1.116827 \n"",
""cv5 1.389543 2.381274 1.491859 \n"",
""\n"",
"" GradientBoostingRegressor MLPRegressor XGBRegressor LGBMRegressor \n"",
""cv1 0.706266 0.670726 0.983154 0.887868 \n"",
""cv2 0.632428 0.837934 0.586317 0.649590 \n"",
""cv3 0.779887 0.874925 0.999408 0.879154 \n"",
""cv4 1.076843 1.366192 1.070108 1.108118 \n"",
""cv5 1.508647 1.533531 1.566166 1.462994 ""
]
},
""execution_count"": 234,
""metadata"": {},
""output_type"": ""execute_result""
}
],
""source"": [
""models=[LinearRegression(),DecisionTreeRegressor(),RandomForestRegressor(),\n"",
"" GradientBoostingRegressor(),\n"",
"" MLPRegressor(hidden_layer_sizes=(50),solver='lbfgs',max_iter=100),\n"",
"" XGBRegressor(n_estimator=100,objective='reg:squarederror'),\n"",
"" LGBMRegressor(n_estimator=100)\n"",
"" ]\n"",
""\n"",
""result=dict()\n"",
""for model in models:\n"",
"" model_name = str(model).split('(')[0]\n"",
"" scores = cross_val_score(model, X=final_feature_train, y=label_train, verbose=0, cv = 5,scoring=make_scorer(mean_squared_error))\n"",
"" result[model_name] = scores\n"",
"" print(model_name + ' is finished')\n"",
""#表格表示\n"",
""result = pd.DataFrame(result)\n"",
""result.index = ['cv' + str(x) for x in range(1, 6)]\n"",
""result""
]
},
{
""cell_type"": ""code"",
""execution_count"": 235,
""id"": ""6f5b9fa1"",
""metadata"": {},
""outputs"": [
{
""data"": {
""text/plain"": [
""LinearRegression 1.189781\n"",
""DecisionTreeRegressor 2.001731\n"",
""RandomForestRegressor 0.947290\n"",
""GradientBoostingRegressor 0.940814\n"",
""MLPRegressor 1.056662\n"",
""XGBRegressor 1.041030\n"",
""LGBMRegressor 0.997545\n"",
""dtype: float64""
]
},
""execution_count"": 235,
""metadata"": {},
""output_type"": ""execute_result""
}
],
""source"": [
""result.mean()""
]
},
{
""cell_type"": ""code"",
""execution_count"": 222,
""id"": ""ab491c21"",
""metadata"": {},
""outputs"": [],
""source"": [
""feature_test=pd.read_csv('Molecular_Descriptor_test.csv',index_col='SMILES')\n"",
""label_test=pd.read_csv('ERα_activity_test.csv',index_col='SMILES')""
]
},
{
""cell_type"": ""code"",
""execution_count"": null,
""id"": ""4cf62651"",
""metadata"": {},
""outputs"": [],
""source"": []
}
],
""metadata"": {
""kernelspec"": {
""display_name"": ""Python 3 (ipykernel)"",
""language"": ""python"",
""name"": ""python3""
},
""language_info"": {
""codemirror_mode"": {
""name"": ""ipython"",
""version"": 3
},
""file_extension"": "".py"",
""mimetype"": ""text/x-python"",
""name"": ""python"",
""nbconvert_exporter"": ""python"",
""pygments_lexer"": ""ipython3"",
""version"": ""3.8.10""
}
},
""nbformat"": 4,
""nbformat_minor"": 5
}
","Unknown"
"ADMET","Bighhhzq/Mathematical-modeling","问题一代码/互信息特征选择.py",".py","2008","38","from sklearn.feature_selection import RFE
from sklearn.ensemble import RandomForestClassifier
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import f1_score,confusion_matrix
from sklearn.metrics import mean_squared_error,mean_absolute_error,mean_absolute_percentage_error,r2_score
import numpy as np
# Create the RFE object and rank each pixel
Set = pd.read_csv('E:\\test\jianmo\特征选择\Clean_Data.csv', encoding='gb18030', index_col=0)
print(Set)
def sigmoid(inx):
return 1.0/(1+np.exp(-inx))
Set_label = Set['pIC50']
print(Set_label)
Set_feature = Set.drop(['SMILES','pIC50'],axis=1)
print(Set_feature)
#Set_feature = Set_feature.loc[:,['ALogP', 'ATSc5', 'nBondsD2', 'nBondsM', 'C2SP3', 'nHBd', 'nHBint2', 'nHBint3', 'nssCH2', 'nsssCH', 'naasC', 'ndsN', 'nssO', 'naaO', 'nssS', 'SdsN', 'SaaO', 'SssS', 'minHBint7', 'minHaaCH', 'mindsN', 'minaaO', 'minssS', 'maxwHBa', 'maxsssCH', 'maxdsN', 'maxaaO', 'ETA_Shape_P', 'ETA_Beta_ns', 'ETA_BetaP_ns', 'ETA_dBeta', 'ETA_Beta_ns_d', 'ETA_BetaP_ns_d', 'nHBDon', 'nHBDon_Lipinski', 'MDEN-23', 'nRing', 'n7Ring', 'nTRing', 'nT7Ring']]
Set_feature = Set_feature.loc[:,['ALogP', 'ATSc1', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'SsOH', 'minHBa', 'minwHBa', 'minHsOH', 'hmin', 'LipoaffinityIndex', 'Kier3', 'MDEC-33', 'WTPT-5','nssCH2', 'nsssCH', 'nssO', 'maxaaCH', 'SHBint10']]
print(Set_feature)
x_train, x_test, y_train, y_test = train_test_split(Set_feature, Set_label , test_size=0.3, random_state=42)
from sklearn.ensemble import RandomForestRegressor #集成学习中的随机森林
rfc = RandomForestRegressor(random_state=42)
rfc = rfc.fit(x_train,y_train)
ac_2 = mean_squared_error(y_test,rfc.predict(x_test))
ridge_mae = mean_absolute_error(y_test, rfc.predict(x_test))
ridge_mape = mean_absolute_percentage_error(y_test, rfc.predict(x_test))
ridge_r2 = r2_score(y_test, rfc.predict(x_test))
print(""MSE ="", ac_2)
print(""MAE ="", ridge_mae)
print(""MAPE ="", ridge_mape)
print(""r2 ="", ridge_r2)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题一代码/递归特征消除RFE.py",".py","1330","33","from sklearn.feature_selection import RFE
from sklearn.ensemble import RandomForestRegressor
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import f1_score,confusion_matrix
from sklearn.metrics import mean_absolute_percentage_error,mean_squared_error,mean_absolute_error,r2_score
# Create the RFE object and rank each pixel
Set = pd.read_csv('E:\\test\jianmo\特征选择\Clean_Data.csv', encoding='gb18030', index_col=0)
print(Set)
Set_label = Set['pIC50']
print(Set_label)
Set_feature = Set.drop(['SMILES','pIC50'],axis=1)
print(Set_feature)
x_train, x_test, y_train, y_test = train_test_split(Set_feature, Set_label , test_size=0.3, random_state=42)
clf_rf_3 = RandomForestRegressor()
rfe = RFE(estimator=clf_rf_3, n_features_to_select=40, step=1)
rfe = rfe.fit(x_train, y_train)
print('Chosen best 25 feature by rfe:',x_train.columns[rfe.support_])
ac = mean_absolute_percentage_error(y_test,rfe.predict(x_test))
print('Accuracy is: ',ac)
ac_2 = mean_squared_error(y_test,rfe.predict(x_test))
ridge_mae = mean_absolute_error(y_test, rfe.predict(x_test))
ridge_mape = mean_absolute_percentage_error(y_test, rfe.predict(x_test))
ridge_r2 = r2_score(y_test, rfe.predict(x_test))
print(""MSE ="", ac_2)
print(""MAE ="", ridge_mae)
print(""MAPE ="", ridge_mape)
print(""r2 ="", ridge_r2)","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题一代码/feature_filter.py",".py","1132","47","import pandas as pd
from 问题一代码.remove_vars import clean_zi_var1
from 问题一代码.dcor import dcor
clean_set0 = pd.read_csv('./clean451.csv', index_col=0, encoding='gb18030')
print(clean_set0)
clean_var_set = clean_set0.drop('SMILES', axis=1)
rps = clean_var_set.corr(method='pearson')
df_ps = rps.iloc[0]
ps_list = rps['pIC50'].tolist()
ps_list.remove(1.0)
feature_set = clean_set0.drop(['SMILES', 'pIC50'], axis=1)
Clean_var = feature_set.columns.tolist()
print(""ps_list:"",ps_list)
print(len(ps_list))
rspm = clean_var_set.corr(method='spearman')
df_spm = rspm.iloc[0]
spm_list = rspm['pIC50'].tolist()
spm_list.remove(1.0)
pic50 = clean_var_set['pIC50'].tolist()
delete2 = []
dcor_list = []
for a in clean_zi_var1:
i = clean_var_set[a].tolist()
d = dcor(pic50, i)
dcor_list.append(d)
if abs(df_ps[a])<0.2 and abs(df_spm[a])<0.2 and d<0.2:
delete2.append(a)
dcor_list.remove(1.0)
print(""dcor_list:"", dcor_list)
df_clean2 = clean_set0.drop(delete2, axis=1)
print(clean_var_set)
print(df_clean2)
Clean_data = df_clean2
print(Clean_data)
Clean_data.to_csv('.\Clean_Data.csv.csv')
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题一代码/dcor.py",".py","954","34","from scipy.spatial.distance import pdist, squareform
import numpy as np
# from numba import jit, float32
def dcor(X, Y):
X = np.atleast_1d(X)
Y = np.atleast_1d(Y)
if np.prod(X.shape) == len(X):
X = X[:, None]
if np.prod(Y.shape) == len(Y):
Y = Y[:, None]
X = np.atleast_2d(X)
Y = np.atleast_2d(Y)
n = X.shape[0]
if Y.shape[0] != X.shape[0]:
raise ValueError('Number of samples must match')
a = squareform(pdist(X))
b = squareform(pdist(Y))
A = a - a.mean(axis=0)[None, :] - a.mean(axis=1)[:, None] + a.mean()
B = b - b.mean(axis=0)[None, :] - b.mean(axis=1)[:, None] + b.mean()
dcov2_xy = (A * B).sum() / float(n * n)
dcov2_xx = (A * A).sum() / float(n * n)
dcov2_yy = (B * B).sum() / float(n * n)
dcor = np.sqrt(dcov2_xy) / np.sqrt(np.sqrt(dcov2_xx) * np.sqrt(dcov2_yy))
return dcor
# a = [1,2,3,4,5]
# b = np.array([1,2,9,4,4])
# d = dcor(a,b)
# print(d)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题一代码/MIC.py",".py","640","26","import pandas as pd
from minepy import MINE
import numpy as np
Set = pd.read_csv('./Clean_Data.csv' , encoding='gb18030', index_col=0)
y_set = Set['pIC50']
feature_set = Set.drop(['SMILES', 'pIC50'], axis=1)
y = np.array(y_set.tolist())
Clean_var = feature_set.columns.tolist()
mine = MINE(alpha=0.6, c=15)
mic = []
for i in Clean_var:
x = np.array(feature_set[i].tolist())
mine.compute_score(x, y)
m = mine.mic()
mic.append(m)
print(mic)
max_index = pd.Series(mic).sort_values().index[:40]
mic_slect_var = [x for x in Clean_var if Clean_var.index(x) in max_index]
print(Clean_var)
print(max_index)
print(mic_slect_var)","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题一代码/集成降维变量选20个.py",".py","3382","71","
import pandas as pd
import heapq
Set = pd.read_csv('E:\\test\jianmo\特征选择\Clean_Data.csv', encoding='gb18030', index_col=0)
print(Set)
Set_label = Set['pIC50']
print(Set_label)
Set_feature = Set.drop(['SMILES','pIC50'],axis=1)
print(Set_feature)
print(Set_feature.columns.tolist())
hu_xx= ['ALogP', 'ATSc5', 'nBondsD2', 'nBondsM', 'C2SP3', 'nHBd', 'nHBint2', 'nHBint3', 'nssCH2', 'nsssCH', 'naasC', 'ndsN', 'nssO', 'naaO', 'nssS', 'SdsN', 'SaaO', 'SssS', 'minHBint7', 'minHaaCH', 'mindsN', 'minaaO', 'minssS', 'maxwHBa', 'maxsssCH', 'maxdsN', 'maxaaO', 'ETA_Shape_P', 'ETA_Beta_ns', 'ETA_BetaP_ns', 'ETA_dBeta', 'ETA_Beta_ns_d', 'ETA_BetaP_ns_d', 'nHBDon', 'nHBDon_Lipinski', 'MDEN-23', 'nRing', 'n7Ring', 'nTRing', 'nT7Ring']
dan_chi2 = ['minHBa', 'ETA_dBeta', 'C1SP2', 'SsOH', 'maxsOH', 'minsOH', 'SsssN',
'minsssN', 'maxsssN', 'SdO', 'maxdO', 'mindO', 'SHBint10', 'minHBint10',
'maxHBint10', 'SssNH', 'nAtomLAC', 'maxssNH', 'minssNH', 'SssO',
'maxssO', 'SaaN', 'minssO', 'nHsOH', 'nHsOH', 'maxaaN', 'minaaN',
'nsssCH', 'SssCH2', 'ndO', 'SHCsats', 'C2SP3', 'nssO', 'ALogP',
'nHBint10', 'C1SP3', 'SHBint4', 'nHCsats', 'nBondsD2', 'nssCH2']
dan_huxinxi = ['SHsOH', 'BCUTc-1l', 'BCUTc-1h', 'minHsOH', 'maxHsOH', 'MLFER_A', 'SHBd', 'Kier3', 'minHBd', 'minHBa', 'hmin', 'ATSc2', 'BCUTp-1h', 'ETA_BetaP_s', 'maxsOH', 'maxHBd', 'ATSc3', 'maxHCsats', 'mindssC', 'minaasC', 'C1SP2', 'minwHBa', 'McGowan_Volume', 'nHBAcc', 'ATSc4', 'VP-6', 'WTPT-5', 'ETA_Alpha', 'SP-2', 'maxssO', 'LipoaffinityIndex',
'SP-6', 'ATSc1', 'minssCH2', 'VPC-6', 'VP-4', 'VPC-5', 'SsOH', 'MAXDP2',
'SP-3']
digui_RFE =['ALogP', 'ATSc1', 'ATSc2', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h','BCUTp-1h', 'C1SP2', 'SC-3', 'VC-5', 'VP-5', 'ECCEN', 'SHBint10', 'SaaCH', 'SsOH', 'minHBa', 'minHBint4', 'minHsOH', 'mindssC', 'minsssN', 'minsOH', 'minssO', 'maxHsOH', 'maxaaCH', 'maxssO', 'gmax', 'hmin','LipoaffinityIndex', 'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'Kier3','MDEC-23', 'MDEC-33', 'MLFER_A', 'MLFER_BH', 'TopoPSA', 'WTPT-5','XLogP']
suiji_senling = ['minsssN', 'MDEC-23', 'LipoaffinityIndex', 'maxHsOH', 'minssO',
'nHBAcc', 'minHsOH', 'BCUTc-1l', 'maxssO', 'MLFER_A', 'C1SP2',
'TopoPSA', 'WTPT-5', 'ATSc3', 'ATSc2', 'VC-5', 'mindssC', 'ATSc1',
'BCUTp-1h', 'BCUTc-1h', 'XLogP', 'MDEC-33', 'Kier3', 'maxsssCH',
'minsOH', 'hmin', 'minHBa', 'SHsOH', 'minHBint7', 'MAXDP2', 'ATSc5',
'maxHBa', 'VCH-7', 'ETA_BetaP_s', 'ATSc4', 'SC-3', 'SsOH', 'minssNH',
'VP-5', 'maxsOH']
l=245
list = [0]*l
for j,i in enumerate(Set_feature.columns.tolist()):
if i in hu_xx:
list[j] = list[j]+1
if i in dan_chi2:
list[j] = list[j]+1
if i in dan_huxinxi:
list[j] = list[j]+1
if i in digui_RFE:
list[j] = list[j]+1
if i in suiji_senling:
list[j] = list[j]+1
print(list)
print(sorted(list,reverse=True)[:60])
result = map(list.index, heapq.nlargest(60, list))
print(result)
best_21_va=[]
for j,i in enumerate(Set_feature.columns.tolist()):
if list[j]>=4:
best_21_va.append(i)
print(best_21_va)
for j,i in enumerate(Set_feature.columns.tolist()):
if list[j]>=3 and list[j]<4:
best_21_va.append(i)
print(best_21_va)
for j,i in enumerate(Set_feature.columns.tolist()):
if list[j]<3 and list[j]>=2:
best_21_va.append(i)
print(best_21_va)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题一代码/基于树的特征选择.py",".py","1877","49","from sklearn.ensemble import RandomForestRegressor
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_percentage_error,mean_squared_error,mean_absolute_error,r2_score
import numpy as np
Set = pd.read_csv('E:\\test\jianmo\特征选择\Clean_Data.csv', encoding='gb18030', index_col=0)
print(Set)
Set_label = Set['pIC50']
print(Set_label)
Set_feature = Set.drop(['SMILES','pIC50'],axis=1)
print(Set_feature)
x_train, x_test, y_train, y_test = train_test_split(Set_feature, Set_label, test_size=0.3, random_state=42)
clf_rf_5 = RandomForestRegressor()
clr_rf_5 = clf_rf_5.fit(x_train,y_train)
ac_2 = mean_squared_error(y_test,clr_rf_5.predict(x_test))
ridge_mae = mean_absolute_error(y_test, clr_rf_5.predict(x_test))
ridge_mape = mean_absolute_percentage_error(y_test, clr_rf_5.predict(x_test))
ridge_r2 = r2_score(y_test, clr_rf_5.predict(x_test))
print(""MSE ="", ac_2)
print(""MAE ="", ridge_mae)
print(""MAPE ="", ridge_mape)
print(""r2 ="", ridge_r2)
importances = clr_rf_5.feature_importances_
std = np.std([tree.feature_importances_ for tree in clr_rf_5.estimators_], axis=0)
indices = np.argsort(importances)[::-1]
# Print the feature ranking
print(""Feature ranking:"")
for f in range(x_train.shape[1]):
print(""%d. feature %d (%f)"" % (f + 1, indices[f], importances[indices[f]]))
# best_index = [136,217,166,146,139,204,126,26,161,221,38,233,240,12,11,47,131,10,28,27,243,218,212,153,137,165,121,103,124,168,14,143,44,186,13,45,115,133,67,159]
# print(Set_feature.columns[best_index])
#
#
# import matplotlib.pyplot as plt
#
# plt.figure(1, figsize=(14, 13))
# plt.title(""Feature importances"")
# plt.bar(range(x_train.shape[1])[:40], importances[best_index], color=""g"" , align=""center"")
# plt.xticks(range(x_train.shape[1])[:40], x_train.columns[best_index],rotation=90)
# plt.xlim([-1, 40])
# plt.show()
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题一代码/remove_vars.py",".py","1531","52","import numpy as np
import pandas as pd
Set = pd.read_csv('./data.csv', encoding='gb18030')
feature = Set.drop('pIC50', axis=1).columns.tolist()
vars_set = Set.drop('SMILES', axis=1)
zi_var = vars_set.drop('pIC50', axis=1).columns.tolist()
print(zi_var)
delete = []
for a in zi_var:
if ((Set[a] == 0).sum() / 1973) > 0.99:
delete.append(a)
clean_Set1 = Set.drop(delete, axis=1)
print(clean_Set1)
clean_Set1.to_csv('./clean451.csv', encoding='gb18030')
clean_zi_set1 = vars_set.drop(delete, axis=1)
clean_zi_var1 = clean_zi_set1.columns.tolist()
def three_sigma(Ser1):
'''
Ser1:表示传入DataFrame的某一列。
'''
rule = (Ser1.mean() - 10 * Ser1.std() > Ser1) | (Ser1.mean() + 10 * Ser1.std() < Ser1)
index = np.arange(Ser1.shape[0])[rule]
return index # 返回落在3sigma之外的行索引值
def delete_out3sigma(Set, var):
""""""
data:待检测的DataFrame
""""""
data1 = Set[var]
data = (data1 - data1.min()) / (data1.max() - data1.min())
out_index = [] # 保存要删除的行索引
for i in range(data.shape[1]): # 对每一列分别用3sigma原则处理
index = three_sigma(data.iloc[:, i])
out_index += index.tolist()
delete_ = list(set(out_index))
print('所删除的行索引为:', delete_)
print(len(delete_))
Set.drop(delete_, inplace=True)
return Set
clean_Set0 = delete_out3sigma(clean_Set1, clean_zi_var1) # 去除异常样本后的结果
clean_Set0.to_csv('./clean_set0.csv', encoding='gb18030')
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题一代码/单变量特征选择chi2.py",".py","1906","48","from sklearn.ensemble import RandomForestRegressor
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_percentage_error,mean_squared_error,mean_absolute_error,r2_score
import numpy as np
Set = pd.read_csv('E:\\test\jianmo\特征选择\Clean_Data.csv', encoding='gb18030', index_col=0)
print(Set)
def sigmoid(inx):
return 1.0/(1+np.exp(-inx))
Set_label = Set['pIC50']
print(Set_label)
Set_feature = Set.drop(['SMILES','pIC50'],axis=1)
print(Set_feature)
Set_feature = sigmoid(Set_feature)
print(Set_feature)
x_train, x_test, y_train, y_test = train_test_split(Set_feature, Set_label.astype(""int"") , test_size=0.3, random_state=42)
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
import heapq
select_feature = SelectKBest(chi2, k=10).fit(x_train, y_train)
print('Score list:', select_feature.scores_)
print(len(select_feature.scores_))
print(sorted(select_feature.scores_,reverse=True)[:10])
result = map(select_feature.scores_.tolist().index, heapq.nlargest(10, select_feature.scores_.tolist()))
# print(list(result))
best_index = list(result)
print(best_index)
print(Set_feature.columns[best_index])
x_train_2 = select_feature.transform(x_train)
x_test_2 = select_feature.transform(x_test)
#random forest classifier with n_estimators=10 (default)
clf_rf_2 = RandomForestRegressor()
clr_rf_2 = clf_rf_2.fit(x_train_2,y_train)
ac_2 = mean_absolute_percentage_error(y_test,clf_rf_2.predict(x_test_2))
print('Accuracy is: ',ac_2)
ac_2 = mean_squared_error(y_test,clf_rf_2.predict(x_test_2))
ridge_mae = mean_absolute_error(y_test, clf_rf_2.predict(x_test_2))
ridge_mape = mean_absolute_percentage_error(y_test, clf_rf_2.predict(x_test_2))
ridge_r2 = r2_score(y_test, clf_rf_2.predict(x_test_2))
print(""MSE ="", ac_2)
print(""MAE ="", ridge_mae)
print(""MAPE ="", ridge_mape)
print(""r2 ="", ridge_r2)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","graph/关系图.py",".py","2200","72","import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
from random import sample
plt.rcParams['pdf.fonttype'] = 42
config = {
""font.family"":'Times New Roman', # 设置字体类型
}
plt.rcParams.update(config)
Set = pd.read_csv('../Clean_Data.csv' , encoding='gb18030', index_col=0)
Set.drop('SMILES', axis=1)
print(Set)
Clean_vars = Set.columns.tolist()
Clean_vars.remove('pIC50')
Cat_vars_low = list(Set.loc[:, (Set.nunique() < 6)].nunique().index)
print(len(Cat_vars_low))
Cat_vars_high = list(Set.loc[:, (Set.nunique() >= 10)].nunique().index)
with plt.rc_context(rc = {'figure.dpi': 600, 'axes.labelsize': 12,
'xtick.labelsize': 12, 'ytick.labelsize': 12}):
fig_3, ax_3 = plt.subplots(3, 3, figsize = (15, 15))
for idx, (column, axes) in list(enumerate(zip(Cat_vars_low[0:9], ax_3.flatten()))):
order = Set.groupby(column)['pIC50'].mean().sort_values(ascending = True).index
plt.xticks(rotation=90)
sns.violinplot(ax = axes, x = Set[column],
y = Set['pIC50'],
order = order, scale = 'width',
linewidth = 0.5, palette = 'viridis',
inner = None)
plt.setp(axes.collections, alpha = 0.3)
sns.stripplot(ax = axes, x = Set[column],
y = Set['pIC50'],
palette = 'viridis', s = 1.5, alpha = 0.75,
order = order, jitter = 0.07)
sns.pointplot(ax = axes, x = Set[column],
y = Set['pIC50'],
order = order,
color = '#ff5736', scale = 0.2,
estimator = np.mean, ci = 'sd',
errwidth = 0.5, capsize = 0.15, join = True)
plt.setp(axes.lines, zorder = 100)
plt.setp(axes.collections, zorder = 100)
if Set[column].nunique() > 5:
plt.setp(axes.get_xticklabels(), rotation = 90)
else:
[axes.set_visible(False) for axes in ax_3.flatten()[idx + 1:]]
plt.savefig('./问题一小提琴.pdf', dpi=600)
# plt.tight_layout()
# plt.show()
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","graph/Q4变化图.py",".py","1626","41","import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
Set = pd.read_csv('./tu2.csv' , encoding='gb18030')
featuren_all = ['nN', 'ATSc2', 'SCH-7', 'VPC-5', 'SP-6',
'SHaaCH', 'SssCH2', 'SsssCH', 'SssO', 'minHBa',
'mindssC', 'maxsCH3', 'maxsssCH', 'maxssO', 'hmin',
'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y','ETA_BetaP', 'ETA_BetaP_s',
'ETA_EtaP_F', 'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'MLFER_E',
'WTPT-4','BCUTc-1l','BCUTp-1l','VCH-6','SC-5',
'SPC-6','VP-3','SHsOH','SdO','minHsOH',
'maxHother','maxdO','MAXDP2','ETA_EtaP_F_L','MDEC-23',
'MLFER_A','TopoPSA','WTPT-2','BCUTc-1h','BCUTp-1h',
'SHBd','SHother','SsOH','minHBd','minaaCH',
'minaasC','maxHBd','maxwHBa','maxHBint8','maxHsOH',
'LipoaffinityIndex','ETA_Eta_R_L','MDEO-11','minssCH2','apol',
'ATSc1','ATSm3','SCH-6','VCH-7','maxsOH',
'ETA_dEpsilon_D','ETA_Shape_P','ETA_dBetaP','ATSm2','VC-5',
'SsCH3','SaaO','MLFER_S','WPATH','C1SP2',
'ALogP','ATSc3','ATSc5','minsssN','nBondsD2',
'nsssCH']
featuren_pre = ['C1SP2', 'SsOH', 'minHBa', 'maxssO', 'ALogP',
'ATSc1', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h',
'BCUTp-1h', 'mindssC', 'minsssN', 'hmin', 'LipoaffinityIndex',
'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'nBondsD2', 'nsssCH']
df_var_all = Set[featuren_all]
df_var_pre = Set[featuren_pre]
print(df_var_pre.describe())
df_var_pre.to_csv('./pred.csv')","Python"
"ADMET","Bighhhzq/Mathematical-modeling","graph/因变量图.py",".py","1441","51","import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
from random import sample
plt.rcParams['pdf.fonttype'] = 42
config = {
""font.family"":'Times New Roman', # 设置字体类型
""font.size"": 80,
}
plt.rcParams.update(config)
Set = pd.read_csv('../data.csv' , encoding='gb18030')
print(Set)
Set.drop('SMILES', axis=1)
Clean_vars = Set.columns.tolist()
Clean_vars.remove('pIC50')
# Cat_vars.remove('Condition2')
# sns.set_theme(rc = {'grid.linewidth': 0.5,
# 'axes.linewidth': 0.75, 'axes.facecolor': '#fff3e9', 'axes.labelcolor': '#6b1000',
# # 'figure.facecolor': '#f7e7da',
# 'xtick.color': '#6b1000', 'ytick.color': '#6b1000'})
with plt.rc_context(rc={'figure.dpi': 600, 'axes.labelsize': 10,
'xtick.labelsize': 12, 'ytick.labelsize': 12}):
fig_0, ax_0 = plt.subplots(1, 1, figsize=(15, 8))
sns.scatterplot(ax=ax_0, x=list(range(0,1974)),
y=Set['pIC50'],
hue=Set['pIC50'],
alpha=0.7,)
my_y_ticks = np.arange(2, 12, 2)
my_x_ticks = np.arange(0, 2000, 200)
plt.xticks(my_x_ticks)
plt.yticks(my_y_ticks)
# Get rid of legend
ax_0.legend([], [], frameon=False)
# Remove empty figures
# plt.tight_layout()
# plt.show()
plt.savefig('./问题一因变量.pdf', dpi=600)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","graph/拟合图.py",".py","3843","129","import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
# 设置
pd.options.display.notebook_repr_html=False # 表格显示
plt.rcParams['figure.dpi'] = 75 # 图形分辨率
# sns.set_theme(style='darkgrid') # 图形主题
sns.set_theme(style='dark') # 图形主题
plt.rcParams['font.sans-serif'] = ['SimHei'] # 用来正常显示中文标签
plt.rcParams['axes.unicode_minus'] = False # 用来正常显示负号
Set1 = pd.read_csv('./tu2.csv' , encoding='gb18030')
x1 = Set1['SsOH']
y1 = Set1['bianhua']
Set2 = pd.read_csv('./tu2_ALogP.csv' , encoding='gb18030')
x2 = Set2['ALogP']
y2 = Set2['bianhua']
Set3 = pd.read_csv('./tu2_ATSc1.csv' , encoding='gb18030')
x3 = Set3['ATSc1']
y3 = Set3['bianhua']
Set4 = pd.read_csv('./tu2_C1SP2.csv' , encoding='gb18030')
x4 = Set4['C1SP2']
y4 = Set4['bianhua']
Set5 = pd.read_csv('./tu2_maxssO.csv' , encoding='gb18030')
x5 = Set5['maxssO']
y5 = Set5['bianhua']
Set6 = pd.read_csv('./tu2_minHBa.csv' , encoding='gb18030')
x6 = Set6['minHBa']
y6= Set6['bianhua']
# Set = pd.read_csv('../nihe.csv' , encoding='gb18030')
# print(Set)
# y0 = Set[""pIC50""]
# y1 = Set[""REa_9""]
# yq = Set.iloc[0:1976:4]
#
# yq.reset_index(drop=True, inplace=True)
# print(yq)
#
# y0 = yq[""y0""]
# y1 = yq[""y1""]
# x = range(0,len(a))
plt.figure(figsize=[20,30])
plt.subplot(2, 3, 1)
# plt.scatter(x, y, marker='o')
plt.plot(x1, y1, color='g', marker='', label='pIC50', markersize=5, linewidth=""2"")
# plt.plot(x, y1, color='b', marker='', label='预测值', markersize=5, linewidth=""2"")
plt.xlabel('SsOH',fontsize=16)
plt.xticks(x1,())
plt.yticks(fontproperties='Times New Roman', size=16)
plt.xticks(fontproperties='Times New Roman', size=16)
plt.legend(fontsize=15)
# plt.show()
plt.subplot(2, 3, 2)
# plt.scatter(x, y, marker='o')
plt.plot(x2, y2, color='g', marker='', label='pIC50', markersize=5, linewidth=""2"")
# plt.plot(x, y1, color='b', marker='', label='预测值', markersize=5, linewidth=""2"")
plt.xlabel('ALogP',fontsize=16)
plt.xticks(x2,())
plt.yticks(fontproperties='Times New Roman', size=16)
plt.xticks(fontproperties='Times New Roman', size=16)
plt.legend(fontsize=15)
# plt.show()
plt.subplot(2, 3, 3)
# plt.scatter(x, y, marker='o')
plt.plot(x3, y3, color='g', marker='', label='pIC50', markersize=5, linewidth=""2"")
# plt.plot(x, y1, color='b', marker='', label='预测值', markersize=5, linewidth=""2"")
plt.xlabel('ATSc1',fontsize=16)
plt.xticks(x3,())
plt.yticks(fontproperties='Times New Roman', size=16)
plt.xticks(fontproperties='Times New Roman', size=16)
plt.legend(fontsize=15)
# plt.show()
plt.subplot(2, 3, 4)
# plt.scatter(x, y, marker='o')
plt.plot(x4, y4, color='g', marker='', label='pIC50', markersize=5, linewidth=""2"")
# plt.plot(x, y1, color='b', marker='', label='预测值', markersize=5, linewidth=""2"")
plt.xlabel('C1SP2',fontsize=16)
plt.xticks(x4,())
plt.yticks(fontproperties='Times New Roman', size=16)
plt.xticks(fontproperties='Times New Roman', size=16)
plt.legend(fontsize=15)
# plt.show()
plt.subplot(2, 3, 5)
# plt.scatter(x, y, marker='o')
plt.plot(x5, y5, color='g', marker='', label='pIC50', markersize=5, linewidth=""2"")
# plt.plot(x, y1, color='b', marker='', label='预测值', markersize=5, linewidth=""2"")
plt.xlabel('maxssO',fontsize=16)
plt.xticks(x5,())
plt.yticks(fontproperties='Times New Roman', size=16)
plt.xticks(fontproperties='Times New Roman', size=16)
plt.legend(fontsize=15)
# plt.show()
plt.subplot(2, 3, 6)
# plt.scatter(x, y, marker='o')
plt.plot(x6, y6, color='g', marker='', label='pIC50', markersize=5, linewidth=""2"")
# plt.plot(x, y1, color='b', marker='', label='预测值', markersize=5, linewidth=""2"")
plt.xlabel('minHBa',fontsize=16)
plt.xticks(x6,())
plt.yticks(fontproperties='Times New Roman', size=16)
plt.xticks(fontproperties='Times New Roman', size=16)
plt.legend(fontsize=15)
plt.show()","Python"
"ADMET","Bighhhzq/Mathematical-modeling","graph/分布图0.py",".py","1881","67","import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
from random import sample
plt.rcParams['pdf.fonttype'] = 42
config = {
""font.family"":'Times New Roman', # 设置字体类型
}
plt.rcParams.update(config)
Set = pd.read_csv('../Clean_Data.csv' , encoding='gb18030', index_col=0)
Set.drop('SMILES', axis=1)
print(Set)
Clean_vars = Set.columns.tolist()
Clean_vars.remove('pIC50')
Cat_vars_low = list(Set.loc[:, (Set.nunique() < 10)].nunique().index)
print(len(Cat_vars_low))
Cat_vars_high = list(Set.loc[:, (Set.nunique() >= 10)].nunique().index)
with plt.rc_context(rc={'figure.dpi': 200, 'axes.labelsize': 8,
'xtick.labelsize': 6, 'ytick.labelsize': 6,
'legend.fontsize': 6, 'legend.title_fontsize': 6,
'axes.titlesize': 9}):
fig_2, ax_2 = plt.subplots(1, 3, figsize=(8.5, 3.5))
for idx, (column, axes) in list(enumerate(zip(['nssO','nHsOH', 'nHBDon_Lipinski'], ax_2.flatten()))):
sns.kdeplot(ax=axes, x=Set['pIC50'],
hue=Set[column].astype('category'),
common_norm=True,
fill=True, alpha=0.2, palette='viridis',
linewidth=0.6)
axes.set_title(str(column), fontsize=9, fontweight='bold', color='#6b1000')
else:
[axes.set_visible(False) for axes in ax_2.flatten()[idx + 1:]]
# Fixing a legend box for a particulal variable
# ax_2_flat = ax_2.flatten()
#
# legend_3 = ax_2_flat[2].get_legend()
# handles_3 = legend_3.legendHandles
# legend_3.remove()
# ax_2_flat[2].legend(handles_3, Set['HouseStyle'].unique(),
# title='HouseStyle', ncol=2)
plt.tight_layout()
plt.show()
plt.savefig('./问题一分布.pdf', dpi=600)
# plt.tight_layout()
# plt.show()
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","graph/散点图.py",".py","2049","54","import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
from random import sample
plt.rcParams['pdf.fonttype'] = 42
config = {
""font.family"":'Times New Roman', # 设置字体类型
""font.size"": 80,
}
plt.rcParams.update(config)
Set = pd.read_csv('../data.csv' , encoding='gb18030')
print(Set)
Set.drop('SMILES', axis=1)
Slect = ['ALogP', 'ATSc1', 'ATSc4', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'SHsOH', 'SsOH', 'minHBa', 'minwHBa', 'minHsOH', 'minsOH', 'maxHsOH', 'maxssO', 'hmin', 'LipoaffinityIndex', 'Kier3', 'MDEC-33', 'WTPT-3', 'WTPT-5', 'ATSc2', 'ATSc3', 'nBondsD2', 'C2SP3', 'VC-5', 'CrippenLogP', 'nssCH2', 'nsssCH', 'nssO', 'SwHBa', 'SHBint10', 'SaaCH', 'SsssN', 'SssO', 'minsssN', 'maxHBa', 'maxHother', 'maxaaCH', 'maxaasC', 'maxsOH', 'MAXDP2', 'ETA_BetaP_s', 'ETA_dBeta', 'MDEC-23', 'MLFER_A', 'MLFER_BO', 'TopoPSA', 'XLogP']
Clean_vars = Set.columns.tolist()
Clean_vars.remove('pIC50')
# Cat_vars.remove('Condition2')
# sns.set_theme(rc = {'grid.linewidth': 0.5,
# 'axes.linewidth': 0.75, 'axes.facecolor': '#fff3e9', 'axes.labelcolor': '#6b1000',
# # 'figure.facecolor': '#f7e7da',
# 'xtick.color': '#6b1000', 'ytick.color': '#6b1000'})
with plt.rc_context(rc={'figure.dpi': 600, 'axes.labelsize': 10,
'xtick.labelsize': 12, 'ytick.labelsize': 12}):
fig_0, ax_0 = plt.subplots(3, 3, figsize=(12, 10))
for idx, (column, axes) in list(enumerate(zip(Slect[0:9], ax_0.flatten()))):
sns.scatterplot(ax=axes, x=Set[column],
y=Set['pIC50'],
hue=Set['pIC50'],
palette='viridis', alpha=0.7, s=8)
# Get rid of legend
axes.legend([], [], frameon=False)
# Remove empty figures
else:
[axes.set_visible(False) for axes in ax_0.flatten()[idx + 1:]]
# plt.tight_layout()
# plt.show()
plt.savefig('./问题一散点图1.pdf', bbox_inches='tight', dpi=600)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","graph/分布图.py",".py","1922","61","import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
from random import sample
plt.rcParams['pdf.fonttype'] = 42
config = {
""font.family"":'Times New Roman', # 设置字体类型
}
plt.rcParams.update(config)
Set = pd.read_csv('../Clean_Data.csv' , encoding='gb18030', index_col=0)
Set.drop('SMILES', axis=1)
print(Set)
Clean_vars = Set.columns.tolist()
Clean_vars.remove('pIC50')
Cat_vars_low = list(Set.loc[:, (Set.nunique() < 10)].nunique().index)
print(len(Cat_vars_low))
Cat_vars_high = list(Set.loc[:, (Set.nunique() >= 10)].nunique().index)
with plt.rc_context(rc={'figure.dpi': 200, 'axes.labelsize': 8,
'xtick.labelsize': 6, 'ytick.labelsize': 6,
'legend.fontsize': 6, 'legend.title_fontsize': 6,
'axes.titlesize': 9}):
fig_2, ax_2 = plt.subplots(1, 3, figsize=(8.5, 3.5))
with plt.rc_context(rc={'figure.dpi': 200, 'axes.labelsize': 8,
'xtick.labelsize': 6, 'ytick.labelsize': 6,
'legend.fontsize': 6, 'legend.title_fontsize': 6,
'axes.titlesize': 9}):
fig_1, ax_1 = plt.subplots(1, 3, figsize=(8, 3))
for idx, (column, axes) in list(enumerate(zip(['nssO','nHsOH', 'nHBDon_Lipinski'], ax_1.flatten()))):
sns.histplot(ax=axes, x=Set['pIC50'],
hue=Set[column].astype('category'), # multiple = 'stack',
alpha=0.15, palette='viridis',
element='step', linewidth=0.6)
axes.set_title(str(column), fontsize=9, fontweight='bold', color='#6b1000')
else:
[axes.set_visible(False) for axes in ax_1.flatten()[idx + 1:]]
plt.tight_layout()
plt.show()
plt.savefig('./问题一分布.pdf', dpi=600)
# plt.tight_layout()
# plt.show()
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","graph/热图.py",".py","4648","90","import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import matplotlib
from matplotlib import rcParams
# matplotlib.use(""pgf"")
# plt.rcParams['pdf.fonttype'] = 42
# pgf_config = {
# ""font.family"":'serif',
# ""font.size"": 35,
# ""pgf.rcfonts"": False,
# ""text.usetex"": True,
# ""pgf.preamble"": [
# r""\usepackage{unicode-math}"",
# r""\setmainfont{Times New Roman}"",
# r""\usepackage{xeCJK}"",
# r""\setCJKmainfont{SimSun}"",
# ],
# }
# rcParams.update(pgf_config)
# rc = {'axes.unicode_minus': False}
# sns.set(context='notebook', style='ticks', font='SimSon', rc=rc)
# Set = pd.read_csv('../Clean_Data.csv' , encoding='gb18030', index_col=0)
# Slect = ['C1SP2', 'SsOH', 'minHBa', 'maxssO', 'ALogP', 'ATSc1', 'ATSc2', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'minHsOH', 'mindssC', 'minsssN', 'minsOH', 'minssO', 'maxHsOH', 'maxsOH', 'hmin', 'LipoaffinityIndex', 'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'Kier3', 'MLFER_A', 'WTPT-5', 'ATSc4', 'nBondsD2', 'C2SP3', 'SC-3', 'VC-5', 'VP-5', 'nssCH2', 'nsssCH', 'nssO', 'SHBint10', 'SHsOH', 'minHBint7', 'minssNH', 'maxsssCH', 'ETA_dBeta', 'MDEC-23', 'MDEC-33', 'TopoPSA', 'XLogP']
# Slect0 = ['C1SP2', 'SsOH', 'minHBa', 'maxssO', 'ALogP', 'ATSc1', 'ATSc2', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'minHsOH', 'mindssC', 'minsssN', 'minsOH', 'minssO', 'maxHsOH', 'maxsOH', 'hmin', 'LipoaffinityIndex', 'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'Kier3', 'MLFER_A', 'WTPT-5', 'nBondsD2', 'nsssCH']
# Slect00 = ['C1SP2', 'SsOH', 'minHBa', 'maxssO', 'ALogP', 'ATSc1', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'mindssC', 'minsssN', 'hmin', 'LipoaffinityIndex', 'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'nBondsD2', 'nsssCH']
Set = pd.read_csv('../clean451.csv' , encoding='gb18030', index_col=0)
feature11 = ['ATSm2', 'BCUTc-1h', 'SCH-6', 'VC-5', 'SHBd','SsCH3', 'SaaO', 'minHBa', 'hmin',
'LipoaffinityIndex', 'FMF', 'MDEC-23', 'MLFER_S','WPATH']
feature40 = ['ATSc1','ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1l',
'SCH-6', 'SC-5', 'VC-5', 'VP-3', 'nHsOH', 'SHBd', 'SaasC',
'SdO', 'minHBa', 'minHsOH', 'minHother', 'minaasC',
'maxssCH2', 'maxaasC', 'maxdO', 'hmin', 'gmin',
'LipoaffinityIndex', 'MAXDP2', 'ETA_Shape_P',
'FMF', 'MDEC-33', 'MLFER_BO', 'TopoPSA', 'WTPT-2', 'WTPT-4']
feature4 = ['ATSc2', 'BCUTc-1l', 'BCUTp-1l', 'VCH-6', 'SC-5', 'SPC-6', 'VP-3',
'SHsOH', 'SdO', 'minHBa', 'minHsOH',
'maxHother', 'maxdO', 'hmin', 'MAXDP2',
'ETA_dEpsilon_B', 'ETA_Shape_Y', 'ETA_EtaP_F_L', 'MDEC-23', 'MLFER_A',
'TopoPSA', 'WTPT-2', 'WTPT-4']
feature22 =['apol', 'ATSc1', 'ATSm3', 'SCH-6', 'VCH-7', 'SP-6', 'SHBd', 'SHsOH', 'SHaaCH', 'minHBa', 'maxsOH', 'ETA_dEpsilon_D', 'ETA_Shape_P', 'ETA_Shape_Y',
'ETA_BetaP_s', 'ETA_dBetaP']
feature3 = ['apol', 'ATSc2', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'bpol', 'VP-0',
'VP-1', 'VP-2', 'CrippenMR', 'ECCEN', 'SHBd', 'SHother', 'SsOH',
'minHBd', 'minHBa', 'minssCH2', 'minaaCH', 'minaasC', 'maxHBd',
'maxwHBa', 'maxHBint8', 'maxHsOH', 'maxaaCH', 'maxaasC', 'hmin',
'LipoaffinityIndex', 'ETA_dEpsilon_B', 'ETA_Shape_Y', 'ETA_EtaP',
'ETA_EtaP_F', 'ETA_Eta_R_L', 'fragC', 'Kier2', 'Kier3',
'McGowan_Volume', 'MDEO-11', 'WTPT-1', 'WTPT-4', 'WPATH']
feature5 = ['nN', 'ATSc2', 'SCH-7', 'VCH-7', 'VPC-5', 'VPC-6', 'SP-6', 'SHaaCH',
'SssCH2', 'SsssCH', 'SssO', 'minHBa', 'mindssC', 'maxsCH3', 'maxsssCH',
'maxssO', 'hmin', 'ETA_Epsilon_1', 'ETA_Epsilon_2', 'ETA_Epsilon_4',
'ETA_dEpsilon_A', 'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y',
'ETA_BetaP', 'ETA_BetaP_s', 'ETA_EtaP_F', 'ETA_Eta_F_L', 'ETA_EtaP_F_L',
'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'nHBAcc_Lipinski', 'MLFER_BO',
'MLFER_S', 'MLFER_E', 'TopoPSA', 'WTPT-3', 'WTPT-4', 'WTPT-5']
feature55 = ['nN', 'ATSc2', 'SCH-7', 'VPC-5', 'SP-6', 'SHaaCH',
'SssCH2', 'SsssCH', 'SssO', 'minHBa', 'mindssC', 'maxsCH3', 'maxsssCH',
'maxssO', 'hmin', 'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y',
'ETA_BetaP', 'ETA_BetaP_s', 'ETA_EtaP_F',
'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'MLFER_E', 'WTPT-4']
df1 = Set[feature55]
r = df1.corr()
f, ax = plt.subplots(figsize= [40,30])
sns.heatmap(r, ax=ax, vmax=1,vmin=-1,annot=True,
cbar_kws={'label': '相关系数'}, cmap='viridis')
plt.xticks(rotation=90) # 将字体进行旋转
plt.yticks(rotation=360)
# plt.savefig('./问题一待检验热图.pdf', bbox_inches='tight', dpi=600)
# plt.savefig('./feature1.pdf', bbox_inches='tight', dpi=600)
plt.show()","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题二代码/LSTM.py",".py","3174","85","import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
import numpy as np
Set = pd.read_csv('..\问题一代码\Clean_Data.csv', encoding='gb18030', index_col=0)
print(Set)
Set_label = Set['pIC50'].copy()
scaler = StandardScaler()
Set.loc[:, Set.columns != 'SMILES'] = scaler.fit_transform(Set.loc[:, Set.columns != 'SMILES'])
print(Set)
Set_unit_index = ['C1SP2', 'SsOH', 'minHBa', 'maxssO', 'ALogP', 'ATSc1', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'mindssC', 'minsssN', 'hmin', 'LipoaffinityIndex', 'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'nBondsD2', 'nsssCH']
Set_feature = Set.loc[:, Set_unit_index]
print(Set_feature)
print(Set_label)
x_train, x_test, y_train, y_test = train_test_split(Set_feature, Set_label.astype('int') , test_size=0.2, random_state=42)
x_train,y_train = np.array(x_train), np.array(y_train)
x_train = np.reshape(x_train,(x_train.shape[0],x_train.shape[1],1))
from keras.models import Sequential
from keras.layers import Dense, LSTM
from keras.layers import Dropout
model =Sequential()
model.add(LSTM(64,return_sequences=True, input_shape=(x_train.shape[1],1)))
model.add(Dropout(0.5))
model.add(LSTM(64, return_sequences= False))
model.add(Dropout(0.5))
model.add(Dense(128))
model.add(Dropout(0.5))
model.add(Dense(64))
model.add(Dropout(0.5))
model.add(Dense(1))
model.summary()
from sklearn.metrics import mean_absolute_error,mean_absolute_percentage_error, mean_squared_error , r2_score
import tensorflow as tf
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=1e-3), loss='mean_squared_error')
x_test = np.array(x_test)
x_test = np.reshape(x_test, (x_test.shape[0], x_test.shape[1],1))
model.fit(x_train,y_train, batch_size=85, epochs=300, validation_data=(x_test,y_test),validation_batch_size=100)
x_test = np.array(x_test)
x_test = np.reshape(x_test, (x_test.shape[0], x_test.shape[1],1))
predictions = model.predict(x_test)
print(predictions)
print(predictions.shape)
print(y_test.shape)
ridge_mae = mean_absolute_error(y_test, predictions)
ridge_mape = mean_absolute_percentage_error(y_test, predictions)
ridge_mse = mean_squared_error(y_test, predictions)
ridge_r2 = r2_score(y_test, predictions)
print(""Ridge MAE ="", ridge_mae)
print(""Ridge MAPE ="", ridge_mape)
print(""Ridge MSE ="", ridge_mse)
print(""Ridge r2 ="", ridge_r2)
Set = pd.read_csv('E:\\test\jianmo\预测\Molecular_test.csv', encoding='gb18030', index_col=0)
print(Set)
scaler = StandardScaler()
Set.loc[:, Set.columns != 'SMILES'] = scaler.fit_transform(Set.loc[:, Set.columns != 'SMILES'])
print(Set)
Set_unit_index = ['C1SP2', 'SsOH', 'minHBa', 'maxssO', 'ALogP', 'ATSc1', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'mindssC', 'minsssN', 'hmin', 'LipoaffinityIndex', 'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'nBondsD2', 'nsssCH']
Set_feature = Set.loc[:, Set_unit_index]
X = Set_feature.copy()
x_test = np.array(X)
x_test = np.reshape(x_test, (x_test.shape[0], x_test.shape[1],1))
predictions = model.predict(x_test)
Set_feature['REa_9'] = predictions
Set_feature['REa_6'] = predictions
Set_feature.to_csv('E:\\test\jianmo\Set_feature22.csv', index=False)","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题二代码/stacking算法(包内含8种算法).py",".py","8888","224","import numpy as np
from datetime import datetime
from sklearn.linear_model import ElasticNetCV, LassoCV, RidgeCV
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.svm import SVR
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import RobustScaler
from sklearn.model_selection import KFold, cross_val_score
from mlxtend.regressor import StackingCVRegressor
from xgboost import XGBRegressor
from lightgbm import LGBMRegressor
import pandas as pd
from sklearn.preprocessing import StandardScaler,PolynomialFeatures
from sklearn.ensemble import RandomForestRegressor
Set = pd.read_csv('..\问题一代码\Clean_Data.csv', encoding='gb18030', index_col=0)
print(Set)
Set_label = Set['pIC50'].copy()
scaler = StandardScaler()
Set.loc[:, Set.columns != 'SMILES'] = scaler.fit_transform(Set.loc[:, Set.columns != 'SMILES'])
print(Set)
Set_unit_index = ['C1SP2', 'SsOH', 'minHBa', 'maxssO', 'ALogP', 'ATSc1', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'mindssC', 'minsssN', 'hmin', 'LipoaffinityIndex', 'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'nBondsD2', 'nsssCH']
Set_feature = Set.loc[:, Set_unit_index]
print(Set_feature)
X = Set_feature
y = Set_label
kfolds = KFold(n_splits=5, shuffle=True, random_state=42)
def rmsle2(y, y_pred):
return mean_squared_error(y, y_pred)
def rmsle(y, y_pred):
return mean_squared_error(y, y_pred)
# build our model scoring function
def cv_rmse(model, X=X):
rmse = -cross_val_score(model, X, y, scoring=""neg_mean_squared_error"", cv=kfolds)
return rmse
alphas_alt = [14.5, 14.6, 14.7, 14.8, 14.9, 15, 15.1, 15.2, 15.3, 15.4, 15.5]
alphas2 = [5e-05, 0.0001, 0.0002, 0.0003, 0.0004, 0.0005, 0.0006, 0.0007, 0.0008]
e_alphas = [0.0001, 0.0002, 0.0003, 0.0004, 0.0005, 0.0006, 0.0007]
e_l1ratio = [0.8, 0.85, 0.9, 0.95, 0.99, 1]
ridge = make_pipeline(RobustScaler(),
RidgeCV(alphas=0.1, cv=kfolds))
lasso = make_pipeline(RobustScaler(),
LassoCV(max_iter=1000, alphas=0.0001,
random_state=42, cv=kfolds))
elasticnet = make_pipeline(RobustScaler(),
ElasticNetCV(max_iter=100, alphas=0.0005,
cv=kfolds, l1_ratio=0.8))
svr = make_pipeline(RobustScaler(),
SVR(C=10, coef0=0, degree=1, kernel='rbf', gamma=0.1)) #tiaowan
gbr = GradientBoostingRegressor(n_estimators=300, learning_rate=0.05,
max_depth=4, max_features='sqrt',
min_samples_leaf=15, min_samples_split=10,
loss='huber', random_state=42)
lightgbm = LGBMRegressor(objective='regression',
num_leaves=6,
learning_rate=0.03,
n_estimators=300,
max_bin=200,
bagging_fraction=0.75,
bagging_freq=5,
bagging_seed=7,
feature_fraction=0.2,
feature_fraction_seed=7,
verbose=-1,
# min_data_in_leaf=2,
# min_sum_hessian_in_leaf=11
)
xgboost = XGBRegressor(learning_rate=0.01, n_estimators=346,
max_depth=3, min_child_weight=0,
gamma=0, subsample=0.7,
colsample_bytree=0.7,
objective='reg:linear', nthread=-1,
scale_pos_weight=1, seed=27,
reg_alpha=0.00006)
# stack
stack_gen = StackingCVRegressor(regressors=(ridge, lasso, elasticnet,
gbr, svr, xgboost, lightgbm),
meta_regressor=xgboost,
use_features_in_secondary=True)
print('TEST score on CV')
score = cv_rmse(ridge)
print(""Kernel Ridge score: {:.4f} ({:.4f})\n"".format(score.mean(), score.std()), datetime.now(), )
score = cv_rmse(lasso)
print(""Lasso score: {:.4f} ({:.4f})\n"".format(score.mean(), score.std()), datetime.now(), )
score = cv_rmse(elasticnet)
print(""ElasticNet score: {:.4f} ({:.4f})\n"".format(score.mean(), score.std()), datetime.now(), )
score = cv_rmse(svr)
print(""SVR score: {:.4f} ({:.4f})\n"".format(score.mean(), score.std()), datetime.now(), )
score = cv_rmse(lightgbm)
print(""Lightgbm score: {:.4f} ({:.4f})\n"".format(score.mean(), score.std()), datetime.now(), )
score = cv_rmse(gbr)
print(""GradientBoosting score: {:.4f} ({:.4f})\n"".format(score.mean(), score.std()), datetime.now(), )
score = cv_rmse(xgboost)
print(""Xgboost score: {:.4f} ({:.4f})\n"".format(score.mean(), score.std()), datetime.now(), )
score = cv_rmse(stack_gen)
print(""stack_gen score: {:.4f} ({:.4f})\n"".format(score.mean(), score.std()), datetime.now(), )
print('START Fit')
print(datetime.now(), 'StackingCVRegressor')
stack_gen_model = stack_gen.fit(np.array(X), np.array(y))
print(datetime.now(), 'elasticnet')
elastic_model_full_data = elasticnet.fit(X, y)
print(datetime.now(), 'lasso')
lasso_model_full_data = lasso.fit(X, y)
print(datetime.now(), 'ridge')
ridge_model_full_data = ridge.fit(X, y)
print(datetime.now(), 'svr')
svr_model_full_data = svr.fit(X, y)
print(datetime.now(), 'GradientBoosting')
gbr_model_full_data = gbr.fit(X, y)
print(datetime.now(), 'xgboost')
xgb_model_full_data = xgboost.fit(X, y)
print(datetime.now(), 'lightgbm')
lgb_model_full_data = lightgbm.fit(X, y)
print((stack_gen_model.predict(np.array(X))))
def blend_models_predict(X):
return ((0.02 * elastic_model_full_data.predict(X)) + \
(0.02 * lasso_model_full_data.predict(X)) + \
(0.02 * lasso_model_full_data.predict(X)) + \
(0.14 * svr_model_full_data.predict(X)) + \
(0.15 * gbr_model_full_data.predict(X)) + \
(0.15 * xgb_model_full_data.predict(X)) + \
(0.15 * lgb_model_full_data.predict(X)) + \
(0.35 * stack_gen_model.predict(np.array(X))))
def blend_models_predict(X):
return ((0.15 * rf_model_full_data.predict(X)) + \
(0.15 * svr_model_full_data.predict(X)) + \
(0.15 * gbr_model_full_data.predict(X)) + \
(0.15 * xgb_model_full_data.predict(X)) + \
(0.15 * lgb_model_full_data.predict(X)) + \
(0.25 * stack_gen_model.predict(np.array(X))))
from sklearn.metrics import mean_absolute_percentage_error,mean_squared_error,mean_absolute_error,r2_score
def all_duliang(model,X=X, y=y):
ac_2 = mean_squared_error(y, model.predict(X))
ridge_mae = mean_absolute_error(y, model.predict(X))
ridge_mape = mean_absolute_percentage_error(y, model.predict(X))
ridge_r2 = r2_score(y, model.predict(X))
print(""MSE ="", ac_2)
print(""MAE ="", ridge_mae)
print(""MAPE ="", ridge_mape)
print(""r2 ="", ridge_r2)
def all_duliang2(model,X=X, y=y):
ac_2 = mean_squared_error(y, model(X))
ridge_mae = mean_absolute_error(y, model(X))
ridge_mape = mean_absolute_percentage_error(y, model(X))
ridge_r2 = r2_score(y, model(X))
print(""MSE ="", ac_2)
print(""MAE ="", ridge_mae)
print(""MAPE ="", ridge_mape)
print(""r2 ="", ridge_r2)
print(blend_models_predict(X))
print('RMSLE score on train data:')
print(rmsle(y, blend_models_predict(X)))
print(""elastic_model_full_data:"")
all_duliang(model = elastic_model_full_data , X=X, y =y)
print(""lasso_model_full_data:"")
all_duliang(model = lasso_model_full_data , X=X, y =y)
print(""ridge_model_full_data:"")
all_duliang(model = ridge_model_full_data , X=X, y =y)
print(""rf_model_full_data:"")
all_duliang(model = rf_model_full_data , X=X, y =y)
print(""svr_model_full_data:"")
all_duliang(model = svr_model_full_data , X=X, y =y)
print(""gbr_model_full_data:"")
all_duliang(model = gbr_model_full_data , X=X, y =y)
print(""xgb_model_full_data:"")
all_duliang(model = xgb_model_full_data , X=X, y =y)
print(""lgb_model_full_data:"")
all_duliang(model = lgb_model_full_data , X=X, y =y)
print(""stack_gen_model:"")
all_duliang(model = stack_gen_model , X=X, y =y)
print(""blend_models_9_zhong:"")
all_duliang2(model = blend_models_predict , X=X, y =y)
print(""blend_models_6_zhong:"")
# 预测test上面的样本
Set = pd.read_csv('E:\\test\jianmo\预测\Molecular_test.csv', encoding='gb18030', index_col=0)
scaler = StandardScaler()
Set.loc[:, Set.columns != 'SMILES'] = scaler.fit_transform(Set.loc[:, Set.columns != 'SMILES'])
Set_unit_index = ['C1SP2', 'SsOH', 'minHBa', 'maxssO', 'ALogP', 'ATSc1', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'mindssC', 'minsssN', 'hmin', 'LipoaffinityIndex', 'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'nBondsD2', 'nsssCH']
Set_feature = Set.loc[:, Set_unit_index]
X = Set_feature.copy()
Set_feature['REa_9'] = blend_models_predict(X)
Set_feature.to_csv('.\Set_feature_stacking.csv', index=False)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/DE优化算法.py",".py","17683","455","from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.ensemble import VotingClassifier
from sklearn.pipeline import make_pipeline
from xgboost.sklearn import XGBClassifier
from lightgbm.sklearn import LGBMClassifier
from sklearn.ensemble import RandomForestClassifier
import pandas as pd
gr = pd.read_csv('E:\\test\jianmo\优化\clean451.csv', index_col=0, encoding='gb18030')
# gr = gr.drop('SMILES', axis=1)
# x = gr.iloc[:, 6:].values
Feature_0 = ['C1SP2', 'SsOH', 'minHBa', 'maxssO', 'ALogP',
'ATSc1', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h',
'BCUTp-1h', 'mindssC', 'minsssN', 'hmin', 'LipoaffinityIndex',
'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'nBondsD2', 'nsssCH']
Feature_1 = ['ATSm2', 'BCUTc-1h', 'SCH-6', 'VC-5', 'SHBd',
'SsCH3', 'SaaO', 'minHBa', 'hmin', 'LipoaffinityIndex',
'FMF', 'MDEC-23', 'MLFER_S', 'WPATH']
Feature_2 = ['apol', 'ATSc1', 'ATSm3', 'SCH-6', 'VCH-7',
'SP-6', 'SHBd', 'SHsOH', 'SHaaCH', 'minHBa',
'maxsOH', 'ETA_dEpsilon_D', 'ETA_Shape_P', 'ETA_Shape_Y', 'ETA_BetaP_s',
'ETA_dBetaP']
Feature_3 = ['ATSc2', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'SHBd',
'SHother', 'SsOH', 'minHBd', 'minHBa','minaaCH',
'minaasC', 'maxHBd', 'maxwHBa', 'maxHBint8','maxHsOH',
'hmin', 'LipoaffinityIndex','ETA_dEpsilon_B', 'ETA_Shape_Y','ETA_EtaP_F',
'ETA_Eta_R_L', 'MDEO-11', 'WTPT-4', 'minssCH2']
Feature_4 = ['ATSc2', 'BCUTc-1l', 'BCUTp-1l', 'VCH-6', 'SC-5',
'SPC-6', 'VP-3', 'SHsOH', 'SdO', 'minHBa',
'minHsOH', 'maxHother', 'maxdO', 'hmin', 'MAXDP2',
'ETA_dEpsilon_B', 'ETA_Shape_Y', 'ETA_EtaP_F_L','MDEC-23', 'MLFER_A',
'TopoPSA', 'WTPT-2', 'WTPT-4']
Feature_5 = ['nN', 'ATSc2', 'SCH-7', 'VPC-5', 'SP-6',
'SHaaCH', 'SssCH2', 'SsssCH', 'SssO', 'minHBa',
'mindssC', 'maxsCH3', 'maxsssCH', 'maxssO', 'hmin',
'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y','ETA_BetaP', 'ETA_BetaP_s',
'ETA_EtaP_F', 'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'MLFER_E', 'WTPT-4'] #26
featuren_all = ['nN', 'ATSc2', 'SCH-7', 'VPC-5', 'SP-6',
'SHaaCH', 'SssCH2', 'SsssCH', 'SssO', 'minHBa',
'mindssC', 'maxsCH3', 'maxsssCH', 'maxssO', 'hmin',
'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y','ETA_BetaP', 'ETA_BetaP_s',
'ETA_EtaP_F', 'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'MLFER_E',
'WTPT-4','BCUTc-1l','BCUTp-1l','VCH-6','SC-5',
'SPC-6','VP-3','SHsOH','SdO','minHsOH',
'maxHother','maxdO','MAXDP2','ETA_EtaP_F_L','MDEC-23',
'MLFER_A','TopoPSA','WTPT-2','BCUTc-1h','BCUTp-1h',
'SHBd','SHother','SsOH','minHBd','minaaCH',
'minaasC','maxHBd','maxwHBa','maxHBint8','maxHsOH',
'LipoaffinityIndex','ETA_Eta_R_L','MDEO-11','minssCH2','apol',
'ATSc1','ATSm3','SCH-6','VCH-7','maxsOH',
'ETA_dEpsilon_D','ETA_Shape_P','ETA_dBetaP','ATSm2','VC-5',
'SsCH3','SaaO','MLFER_S','WPATH','C1SP2',
'ALogP','ATSc3','ATSc5','minsssN','nBondsD2',
'nsssCH']
Set = pd.read_csv('E:\\test\jianmo\优化\clean451.csv', index_col=0, encoding='gb18030')
print(Set)
scaler = StandardScaler()
Set.loc[:, Set.columns != 'SMILES'] = scaler.fit_transform(Set.loc[:, Set.columns != 'SMILES'])
Set_d = Set[featuren_all].copy()
print(Set_d)
print(Set_d.shape)
import numpy as np
print(type(Set_d))
print(np.max(Set_d))
print(np.min(Set_d))
max_list=np.max(Set_d).tolist()
min_list=np.min(Set_d).tolist()
print(max_list)
print(min_list)
################################# ""MN""
feature_df = gr[Feature_5].copy()
x = feature_df.values
y_var = ['MN']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=42, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
# pipe1 = make_pipeline(StandardScaler(), clf_lr)
pipe6_1 = make_pipeline(StandardScaler(), rf)
pipe6_2 = make_pipeline(StandardScaler(), xgboost)
pipe6_3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe6_1),
('xgb', pipe6_2),
('lgbm', pipe6_3)
]
ensembel6_4 = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
all_model = [pipe6_1, pipe6_2, pipe6_3, ensembel6_4]
clf_labels = ['RandomForestClassifier', 'XGBClassifier', ""LGBMClassifier"", 'Ensemble']
score = cross_val_score(estimator=pipe6_1, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe6_1.fit(x_train, y_train)
score = cross_val_score(estimator=pipe6_2, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe6_2.fit(x_train, y_train)
score = cross_val_score(estimator=pipe6_3, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe6_3.fit(x_train, y_train)
score = cross_val_score(estimator=ensembel6_4, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
ensembel6_4.fit(x_train, y_train)
################################# ""HOB""
feature_df = gr[Feature_4].copy()
x = feature_df.values
y_var = ['HOB']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=42, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
pipe5_1 = make_pipeline(StandardScaler(), rf)
pipe5_2 = make_pipeline(StandardScaler(), xgboost)
pipe5_3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe5_1),
('xgb', pipe5_2),
('lgbm', pipe5_3)
]
ensembel5_4 = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
all_model = [pipe5_1, pipe5_2, pipe5_3, ensembel5_4]
clf_labels = ['RandomForestClassifier', 'XGBClassifier', ""LGBMClassifier"", 'Ensemble']
score = cross_val_score(estimator=pipe5_1, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe5_1.fit(x_train, y_train)
score = cross_val_score(estimator=pipe5_2, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe5_2.fit(x_train, y_train)
score = cross_val_score(estimator=pipe5_3, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe5_3.fit(x_train, y_train)
score = cross_val_score(estimator=ensembel5_4, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
ensembel5_4.fit(x_train, y_train)
################################# ""hERG""
feature_df = gr[Feature_3].copy()
x = feature_df.values
y_var = ['hERG']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=42, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
pipe4_1 = make_pipeline(StandardScaler(), rf)
pipe4_2 = make_pipeline(StandardScaler(), xgboost)
pipe4_3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe4_1),
('xgb', pipe4_2),
('lgbm', pipe4_3)
]
ensembel4_4 = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
all_model = [pipe4_1, pipe4_2, pipe4_3, ensembel4_4]
clf_labels = ['RandomForestClassifier', 'XGBClassifier', ""LGBMClassifier"", 'Ensemble']
score = cross_val_score(estimator=pipe4_1, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe4_1.fit(x_train, y_train)
score = cross_val_score(estimator=pipe4_2, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe4_2.fit(x_train, y_train)
score = cross_val_score(estimator=pipe4_3, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe4_3.fit(x_train, y_train)
score = cross_val_score(estimator=ensembel4_4, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
ensembel4_4.fit(x_train, y_train)
################################# ""CYP3A4""
feature_df = gr[Feature_2].copy()
x = feature_df.values
y_var = ['CYP3A4']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=42, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
pipe3_1 = make_pipeline(StandardScaler(), rf)
pipe3_2 = make_pipeline(StandardScaler(), xgboost)
pipe3_3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe3_1),
('xgb', pipe3_2),
('lgbm', pipe3_3)
]
ensembel3_4 = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
all_model = [pipe3_1, pipe3_2, pipe3_3, ensembel3_4]
clf_labels = ['RandomForestClassifier', 'XGBClassifier', ""LGBMClassifier"", 'Ensemble']
score = cross_val_score(estimator=pipe3_1, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe3_1.fit(x_train, y_train)
score = cross_val_score(estimator=pipe3_2, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe3_2.fit(x_train, y_train)
score = cross_val_score(estimator=pipe3_3, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe3_3.fit(x_train, y_train)
score = cross_val_score(estimator=ensembel3_4, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
ensembel3_4.fit(x_train, y_train)
################################# ""Caco-2""
feature_df = gr[Feature_1].copy()
x = feature_df.values
y_var = ['Caco-2']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=42, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
pipe2_1 = make_pipeline(StandardScaler(), rf)
pipe2_2 = make_pipeline(StandardScaler(), xgboost)
pipe2_3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe2_1),
('xgb', pipe2_2),
('lgbm', pipe2_3)
]
ensembel2_4 = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
score = cross_val_score(estimator=pipe2_1, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe2_1.fit(x_train, y_train)
score = cross_val_score(estimator=pipe2_2, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe2_2.fit(x_train, y_train)
score = cross_val_score(estimator=pipe2_3, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe2_3.fit(x_train, y_train)
score = cross_val_score(estimator=ensembel2_4, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
ensembel2_4.fit(x_train, y_train)
################################# ""ERa""
from xgboost import XGBRegressor
from sklearn.ensemble import GradientBoostingRegressor
from mlxtend.regressor import StackingCVRegressor
Set = pd.read_csv('E:\\test\jianmo\预测\Clean_Data.csv', encoding='gb18030', index_col=0)
y_train = Set['pIC50'].copy()
Set.loc[:, Set.columns != 'SMILES'] = scaler.fit_transform(Set.loc[:, Set.columns != 'SMILES'])
x_train = Set[Feature_0].copy()
x_train = x_train.values
print('训练集和测试集 shape', x_train.shape, y_train.shape)
gbr = GradientBoostingRegressor(n_estimators=3000, learning_rate=0.05,
max_depth=4, max_features='sqrt',
min_samples_leaf=15, min_samples_split=10,
loss='huber', random_state=42)
xgboost = XGBRegressor(learning_rate=0.01, n_estimators=3460,
max_depth=3, min_child_weight=0,
gamma=0, subsample=0.7,
colsample_bytree=0.7,
objective='reg:linear', nthread=-1,
scale_pos_weight=1, seed=27,
reg_alpha=0.00006)
stack_gen = StackingCVRegressor(regressors=( gbr, xgboost), meta_regressor=xgboost, use_features_in_secondary=True)
pipe1_1 = make_pipeline(StandardScaler(), stack_gen)
from sklearn.model_selection import cross_val_score
score = cross_val_score(estimator=pipe1_1, X=x_train, y=y_train, cv=10, scoring='neg_mean_squared_error')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe1_1.fit(x_train, y_train)
def schaffer(p):
'''
This function has plenty of local minimum, with strong shocks
global minimum at (0,0) with value 0
'''
x1, x2, x3, x4, x5, x6, x7, x8, x9, x10, x11, x12, x13, x14, x15, x16, x17, x18, x19, x20, x21, x22, x23, x24, x25, x26\
,x27,x28,x29,x30,x31,x32,x33,x34,x35,x36,x37,x38,x39,x40,x41,x42,x43 \
,x44,x45,x46,x47,x48,x49,x50,x51,x52,x53,x54,x55,x56,x57,x58,x59\
,x60,x61,x62,x63,x64,x65,x66,x67,x68\
,x69,x70,x71,x72,x73,x74\
,x75,x76,x77,x78,x79,x80,x81= p
feature1 = [x1, x2, x3, x4, x5, x6, x7, x8, x9, x10, x11, x12, x13, x14, x15, x16, x17, x18, x19, x20, x21, x22, x23, x24, x25, x26] # MN
feature2 = [x2,x27,x28,x29,x30,x31,x32,x33,x34,x10,x35,x36,x37,x15,x38,x16,x18,x39,x40,x41,x42,x43,x26] # Hob
feature3 = [x2,x27,x44,x45,x46,x47,x48,x49,x10,x50,x51,x52,x53,x54,x55,x15,x56,x16,x18,x21,x57,x58,x26,x59] #hERG
feature4 = [x60,x61,x62,x63,x64,x5,x46,x33,x6,x10,x65,x66,x67,x18,x20,x68] #CYP3A4
feature5 = [x69,x44,x63,x70,x46,x71,x72,x10,x15,x56,x23,x40,x73,x74]
feature6 = [x75,x48,x10,x14,x76,x61,x77,x78,x27,x44,x45,x45,x79,x45,x56,x38,x20,x24,x80,x81]
pipe6_1_score = pipe6_1.predict([feature1])
pipe6_2_score = pipe6_2.predict([feature1])
pipe6_3_score = pipe6_3.predict([feature1])
pipe6_4_score = ensembel6_4.predict([feature1])
score_mn =pipe6_1_score + pipe6_2_score + pipe6_3_score + pipe6_4_score
pipe5_1_score = pipe5_1.predict([feature2])
pipe5_2_score = pipe5_2.predict([feature2])
pipe5_3_score = pipe5_3.predict([feature2])
pipe5_4_score = ensembel5_4.predict([feature2])
score_hob =pipe5_1_score + pipe5_2_score + pipe5_3_score + pipe5_4_score
pipe4_1_score = pipe4_1.predict([feature3])
pipe4_2_score = pipe4_2.predict([feature3])
pipe4_3_score = pipe4_3.predict([feature3])
pipe4_4_score = ensembel4_4.predict([feature3])
score_hERG =pipe4_1_score + pipe4_2_score + pipe4_3_score + pipe4_4_score
pipe3_1_score = pipe3_1.predict([feature4])
pipe3_2_score = pipe3_2.predict([feature4])
pipe3_3_score = pipe3_3.predict([feature4])
pipe3_4_score = ensembel3_4.predict([feature4])
score_CYP3A4 =pipe3_1_score + pipe3_2_score + pipe3_3_score + pipe3_4_score
pipe2_1_score = pipe2_1.predict([feature5])
pipe2_2_score = pipe2_2.predict([feature5])
pipe2_3_score = pipe2_3.predict([feature5])
pipe2_4_score = ensembel2_4.predict([feature5])
score_Caco2 =pipe2_1_score + pipe2_2_score + pipe2_3_score + pipe2_4_score
pipe1_1_score = pipe1_1.predict([feature6])
score_ERa =pipe1_1_score
print(score_ERa)
print(score_Caco2)
print(score_CYP3A4)
print(score_hERG)
print(score_hob)
print(score_mn)
print(""总得分"", -score_Caco2 + score_CYP3A4 + score_hERG - score_hob + score_mn - score_ERa)
return -score_Caco2 + score_CYP3A4 + score_hERG - score_hob + score_mn - score_ERa
from 问题四优化.scikitopt.sko.DE import DE
de = DE(func=schaffer, n_dim=81, size_pop=100, max_iter=40, lb=min_list, ub=max_list)
# pso.run()
best_x, best_y = de.run()
print('best_x:', best_x, '\n', 'best_y:', best_y)
import matplotlib.pyplot as plt
plt.plot(de.gbest_y_hist)
plt.show()
import pandas as pd
import matplotlib.pyplot as plt
Y_history = pd.DataFrame(de.all_history_Y)
fig, ax = plt.subplots(2, 1)
ax[0].plot(Y_history.index, Y_history.values, '.', color='red')
Y_history.min(axis=1).cummin().plot(kind='line')
plt.show()
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/PSO优化算法.py",".py","17926","453","from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.ensemble import VotingClassifier
from sklearn.pipeline import make_pipeline
from xgboost.sklearn import XGBClassifier
from lightgbm.sklearn import LGBMClassifier
from sklearn.ensemble import RandomForestClassifier
import pandas as pd
gr = pd.read_csv('E:\\test\jianmo\优化\clean451.csv', index_col=0, encoding='gb18030')
# gr = gr.drop('SMILES', axis=1)
# x = gr.iloc[:, 6:].values
Feature_0 = ['C1SP2', 'SsOH', 'minHBa', 'maxssO', 'ALogP',
'ATSc1', 'ATSc3', 'ATSc5', 'BCUTc-1l', 'BCUTc-1h',
'BCUTp-1h', 'mindssC', 'minsssN', 'hmin', 'LipoaffinityIndex',
'MAXDP2', 'ETA_BetaP_s', 'nHBAcc', 'nBondsD2', 'nsssCH'] #问题二变量
Feature_1 = ['ATSm2', 'BCUTc-1h', 'SCH-6', 'VC-5', 'SHBd',
'SsCH3', 'SaaO', 'minHBa', 'hmin', 'LipoaffinityIndex',
'FMF', 'MDEC-23', 'MLFER_S', 'WPATH'] #问题三_第一类分类变量
Feature_2 = ['apol', 'ATSc1', 'ATSm3', 'SCH-6', 'VCH-7',
'SP-6', 'SHBd', 'SHsOH', 'SHaaCH', 'minHBa',
'maxsOH', 'ETA_dEpsilon_D', 'ETA_Shape_P', 'ETA_Shape_Y', 'ETA_BetaP_s',
'ETA_dBetaP'] #问题三_第二类分类变量
Feature_3 = ['ATSc2', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'SHBd',
'SHother', 'SsOH', 'minHBd', 'minHBa','minaaCH',
'minaasC', 'maxHBd', 'maxwHBa', 'maxHBint8','maxHsOH',
'hmin', 'LipoaffinityIndex','ETA_dEpsilon_B', 'ETA_Shape_Y','ETA_EtaP_F',
'ETA_Eta_R_L', 'MDEO-11', 'WTPT-4', 'minssCH2'] #问题三_第三类分类变量
Feature_4 = ['ATSc2', 'BCUTc-1l', 'BCUTp-1l', 'VCH-6', 'SC-5',
'SPC-6', 'VP-3', 'SHsOH', 'SdO', 'minHBa',
'minHsOH', 'maxHother', 'maxdO', 'hmin', 'MAXDP2',
'ETA_dEpsilon_B', 'ETA_Shape_Y', 'ETA_EtaP_F_L','MDEC-23', 'MLFER_A',
'TopoPSA', 'WTPT-2', 'WTPT-4'] #问题三_第四类分类变量
Feature_5 = ['nN', 'ATSc2', 'SCH-7', 'VPC-5', 'SP-6',
'SHaaCH', 'SssCH2', 'SsssCH', 'SssO', 'minHBa',
'mindssC', 'maxsCH3', 'maxsssCH', 'maxssO', 'hmin',
'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y','ETA_BetaP', 'ETA_BetaP_s',
'ETA_EtaP_F', 'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'MLFER_E', 'WTPT-4'] #问题四_第四类分类变量
featuren_all = ['nN', 'ATSc2', 'SCH-7', 'VPC-5', 'SP-6',
'SHaaCH', 'SssCH2', 'SsssCH', 'SssO', 'minHBa',
'mindssC', 'maxsCH3', 'maxsssCH', 'maxssO', 'hmin',
'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y','ETA_BetaP', 'ETA_BetaP_s',
'ETA_EtaP_F', 'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'MLFER_E',
'WTPT-4','BCUTc-1l','BCUTp-1l','VCH-6','SC-5',
'SPC-6','VP-3','SHsOH','SdO','minHsOH',
'maxHother','maxdO','MAXDP2','ETA_EtaP_F_L','MDEC-23',
'MLFER_A','TopoPSA','WTPT-2','BCUTc-1h','BCUTp-1h',
'SHBd','SHother','SsOH','minHBd','minaaCH',
'minaasC','maxHBd','maxwHBa','maxHBint8','maxHsOH',
'LipoaffinityIndex','ETA_Eta_R_L','MDEO-11','minssCH2','apol',
'ATSc1','ATSm3','SCH-6','VCH-7','maxsOH',
'ETA_dEpsilon_D','ETA_Shape_P','ETA_dBetaP','ATSm2','VC-5',
'SsCH3','SaaO','MLFER_S','WPATH','C1SP2',
'ALogP','ATSc3','ATSc5','minsssN','nBondsD2',
'nsssCH'] # 汇总的的第四问变量81个
Set = pd.read_csv('E:\\test\jianmo\优化\clean451.csv', index_col=0, encoding='gb18030')
print(Set)
scaler = StandardScaler()
Set.loc[:, Set.columns != 'SMILES'] = scaler.fit_transform(Set.loc[:, Set.columns != 'SMILES'])
Set_d = Set[featuren_all].copy()
print(Set_d)
print(Set_d.shape)
import numpy as np
print(type(Set_d))
print(np.max(Set_d))
print(np.min(Set_d))
max_list=np.max(Set_d).tolist()
min_list=np.min(Set_d).tolist()
print(max_list)
print(min_list)
################################# ""MN""
feature_df = gr[Feature_5].copy()
x = feature_df.values
y_var = ['MN']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=42, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
pipe6_1 = make_pipeline(StandardScaler(), rf)
pipe6_2 = make_pipeline(StandardScaler(), xgboost)
pipe6_3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe6_1),
('xgb', pipe6_2),
('lgbm', pipe6_3)
]
ensembel6_4 = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
all_model = [pipe6_1, pipe6_2, pipe6_3, ensembel6_4]
clf_labels = ['RandomForestClassifier', 'XGBClassifier', ""LGBMClassifier"", 'Ensemble']
score = cross_val_score(estimator=pipe6_1, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe6_1.fit(x_train, y_train)
score = cross_val_score(estimator=pipe6_2, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe6_2.fit(x_train, y_train)
score = cross_val_score(estimator=pipe6_3, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe6_3.fit(x_train, y_train)
score = cross_val_score(estimator=ensembel6_4, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
ensembel6_4.fit(x_train, y_train)
################################# ""HOB""
feature_df = gr[Feature_4].copy()
x = feature_df.values
y_var = ['HOB']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=42, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
pipe5_1 = make_pipeline(StandardScaler(), rf)
pipe5_2 = make_pipeline(StandardScaler(), xgboost)
pipe5_3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe5_1),
('xgb', pipe5_2),
('lgbm', pipe5_3)
]
ensembel5_4 = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
all_model = [pipe5_1, pipe5_2, pipe5_3, ensembel5_4]
clf_labels = ['RandomForestClassifier', 'XGBClassifier', ""LGBMClassifier"", 'Ensemble']
score = cross_val_score(estimator=pipe5_1, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe5_1.fit(x_train, y_train)
score = cross_val_score(estimator=pipe5_2, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe5_2.fit(x_train, y_train)
score = cross_val_score(estimator=pipe5_3, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe5_3.fit(x_train, y_train)
score = cross_val_score(estimator=ensembel5_4, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
ensembel5_4.fit(x_train, y_train)
################################# ""hERG""
feature_df = gr[Feature_3].copy()
x = feature_df.values
y_var = ['hERG']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=42, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
pipe4_1 = make_pipeline(StandardScaler(), rf)
pipe4_2 = make_pipeline(StandardScaler(), xgboost)
pipe4_3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe4_1),
('xgb', pipe4_2),
('lgbm', pipe4_3)
]
ensembel4_4 = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
all_model = [pipe4_1, pipe4_2, pipe4_3, ensembel4_4]
clf_labels = ['RandomForestClassifier', 'XGBClassifier', ""LGBMClassifier"", 'Ensemble']
score = cross_val_score(estimator=pipe4_1, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe4_1.fit(x_train, y_train)
score = cross_val_score(estimator=pipe4_2, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe4_2.fit(x_train, y_train)
score = cross_val_score(estimator=pipe4_3, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe4_3.fit(x_train, y_train)
score = cross_val_score(estimator=ensembel4_4, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
ensembel4_4.fit(x_train, y_train)
################################# ""CYP3A4""
feature_df = gr[Feature_2].copy()
x = feature_df.values
y_var = ['CYP3A4']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=42, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
pipe3_1 = make_pipeline(StandardScaler(), rf)
pipe3_2 = make_pipeline(StandardScaler(), xgboost)
pipe3_3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe3_1),
('xgb', pipe3_2),
('lgbm', pipe3_3)
]
ensembel3_4 = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
all_model = [pipe3_1, pipe3_2, pipe3_3, ensembel3_4]
clf_labels = ['RandomForestClassifier', 'XGBClassifier', ""LGBMClassifier"", 'Ensemble']
score = cross_val_score(estimator=pipe3_1, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe3_1.fit(x_train, y_train)
score = cross_val_score(estimator=pipe3_2, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe3_2.fit(x_train, y_train)
score = cross_val_score(estimator=pipe3_3, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe3_3.fit(x_train, y_train)
score = cross_val_score(estimator=ensembel3_4, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
ensembel3_4.fit(x_train, y_train)
################################# ""Caco-2""
feature_df = gr[Feature_1].copy()
x = feature_df.values
y_var = ['Caco-2']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=42, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
pipe2_1 = make_pipeline(StandardScaler(), rf)
pipe2_2 = make_pipeline(StandardScaler(), xgboost)
pipe2_3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe2_1),
('xgb', pipe2_2),
('lgbm', pipe2_3)
]
ensembel2_4 = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
score = cross_val_score(estimator=pipe2_1, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe2_1.fit(x_train, y_train)
score = cross_val_score(estimator=pipe2_2, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe2_2.fit(x_train, y_train)
score = cross_val_score(estimator=pipe2_3, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe2_3.fit(x_train, y_train)
score = cross_val_score(estimator=ensembel2_4, X=x_train, y=y_train, cv=10, scoring='roc_auc')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
ensembel2_4.fit(x_train, y_train)
################################# ""ERa""
from xgboost import XGBRegressor
from sklearn.ensemble import GradientBoostingRegressor
from mlxtend.regressor import StackingCVRegressor
Set = pd.read_csv('E:\\test\jianmo\预测\Clean_Data.csv', encoding='gb18030', index_col=0)
y_train = Set['pIC50'].copy()
Set.loc[:, Set.columns != 'SMILES'] = scaler.fit_transform(Set.loc[:, Set.columns != 'SMILES'])
x_train = Set[Feature_0].copy()
x_train = x_train.values
print('训练集和测试集 shape', x_train.shape, y_train.shape)
gbr = GradientBoostingRegressor(n_estimators=3000, learning_rate=0.05,
max_depth=4, max_features='sqrt',
min_samples_leaf=15, min_samples_split=10,
loss='huber', random_state=42)
xgboost = XGBRegressor(learning_rate=0.01, n_estimators=3460,
max_depth=3, min_child_weight=0,
gamma=0, subsample=0.7,
colsample_bytree=0.7,
objective='reg:linear', nthread=-1,
scale_pos_weight=1, seed=27,
reg_alpha=0.00006)
stack_gen = StackingCVRegressor(regressors=( gbr, xgboost), meta_regressor=xgboost, use_features_in_secondary=True)
pipe1_1 = make_pipeline(StandardScaler(), stack_gen)
from sklearn.model_selection import cross_val_score
score = cross_val_score(estimator=pipe1_1, X=x_train, y=y_train, cv=10, scoring='neg_mean_squared_error')
print(""ElasticNet score: {:.4f}\n"".format(score.mean(), score.std()), )
pipe1_1.fit(x_train, y_train)
# 优化目标函数schaffer的构建---------------------------------------------------------
def schaffer(p):
x1, x2, x3, x4, x5, x6, x7, x8, x9, x10, x11, x12, x13, x14, x15, x16, x17, x18, x19, x20, x21, x22, x23, x24, x25, x26\
,x27,x28,x29,x30,x31,x32,x33,x34,x35,x36,x37,x38,x39,x40,x41,x42,x43 \
,x44,x45,x46,x47,x48,x49,x50,x51,x52,x53,x54,x55,x56,x57,x58,x59\
,x60,x61,x62,x63,x64,x65,x66,x67,x68\
,x69,x70,x71,x72,x73,x74\
,x75,x76,x77,x78,x79,x80,x81= p
feature1 = [x1, x2, x3, x4, x5, x6, x7, x8, x9, x10, x11, x12, x13, x14, x15, x16, x17, x18, x19, x20, x21, x22, x23, x24, x25, x26] # MN
feature2 = [x2,x27,x28,x29,x30,x31,x32,x33,x34,x10,x35,x36,x37,x15,x38,x16,x18,x39,x40,x41,x42,x43,x26] # Hob
feature3 = [x2,x27,x44,x45,x46,x47,x48,x49,x10,x50,x51,x52,x53,x54,x55,x15,x56,x16,x18,x21,x57,x58,x26,x59] #hERG
feature4 = [x60,x61,x62,x63,x64,x5,x46,x33,x6,x10,x65,x66,x67,x18,x20,x68] #CYP3A4
feature5 = [x69,x44,x63,x70,x46,x71,x72,x10,x15,x56,x23,x40,x73,x74]
feature6 = [x75,x48,x10,x14,x76,x61,x77,x78,x27,x44,x45,x45,x79,x45,x56,x38,x20,x24,x80,x81]
pipe6_1_score = pipe6_1.predict([feature1])
pipe6_2_score = pipe6_2.predict([feature1])
pipe6_3_score = pipe6_3.predict([feature1])
pipe6_4_score = ensembel6_4.predict([feature1])
score_mn =pipe6_1_score + pipe6_2_score + pipe6_3_score + pipe6_4_score
pipe5_1_score = pipe5_1.predict([feature2])
pipe5_2_score = pipe5_2.predict([feature2])
pipe5_3_score = pipe5_3.predict([feature2])
pipe5_4_score = ensembel5_4.predict([feature2])
score_hob =pipe5_1_score + pipe5_2_score + pipe5_3_score + pipe5_4_score
pipe4_1_score = pipe4_1.predict([feature3])
pipe4_2_score = pipe4_2.predict([feature3])
pipe4_3_score = pipe4_3.predict([feature3])
pipe4_4_score = ensembel4_4.predict([feature3])
score_hERG =pipe4_1_score + pipe4_2_score + pipe4_3_score + pipe4_4_score
pipe3_1_score = pipe3_1.predict([feature4])
pipe3_2_score = pipe3_2.predict([feature4])
pipe3_3_score = pipe3_3.predict([feature4])
pipe3_4_score = ensembel3_4.predict([feature4])
score_CYP3A4 =pipe3_1_score + pipe3_2_score + pipe3_3_score + pipe3_4_score
pipe2_1_score = pipe2_1.predict([feature5])
pipe2_2_score = pipe2_2.predict([feature5])
pipe2_3_score = pipe2_3.predict([feature5])
pipe2_4_score = ensembel2_4.predict([feature5])
score_Caco2 =pipe2_1_score + pipe2_2_score + pipe2_3_score + pipe2_4_score
pipe1_1_score = pipe1_1.predict([feature6])
score_ERa =pipe1_1_score
print(score_ERa)
print(score_Caco2)
print(score_CYP3A4)
print(score_hERG)
print(score_hob)
print(score_mn)
print(""总得分"", -score_Caco2 + score_CYP3A4 + score_hERG - score_hob + score_mn - score_ERa)
return -score_Caco2 + score_CYP3A4 + score_hERG - score_hob + score_mn - score_ERa
from ..scikitopt.sko.PSO import PSO
pso = PSO(func=schaffer, dim=81, pop=300, max_iter=80, lb=min_list, ub=max_list, w=0.8, c1=0.5, c2=0.5)
pso = PSO(func=schaffer, dim=81, pop=20, max_iter=20, lb=min_list, ub=max_list, w=0.8, c1=0.5, c2=0.5)
best_x, best_y = pso.run()
print('best_x:', best_x, '\n', 'best_y:', best_y)
import matplotlib.pyplot as plt
plt.plot(pso.gbest_y_hist)
plt.show()
import pandas as pd
import matplotlib.pyplot as plt
Y_history = pd.DataFrame(pso.all_history_Y)
fig, ax = plt.subplots(2, 1)
ax[0].plot(Y_history.index, Y_history.values, '.', color='red')
Y_history.min(axis=1).cummin().plot(kind='line')
plt.show()
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/最优解_最近的样本.py",".py","4938","130","import pandas as pd
from sklearn.preprocessing import StandardScaler
Set = pd.read_csv('E:\\test\jianmo\优化\clean451.csv', encoding='gb18030', index_col=0)
print(Set)
scaler = StandardScaler()
Set2 = Set.copy()
print(Set['nN'])
Set.loc[:, Set.columns != 'SMILES'] = scaler.fit_transform(Set.loc[:, Set.columns != 'SMILES'])
print(Set)
featuren_all = ['nN', 'ATSc2', 'SCH-7', 'VPC-5', 'SP-6',
'SHaaCH', 'SssCH2', 'SsssCH', 'SssO', 'minHBa',
'mindssC', 'maxsCH3', 'maxsssCH', 'maxssO', 'hmin',
'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y','ETA_BetaP', 'ETA_BetaP_s',
'ETA_EtaP_F', 'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'MLFER_E',
'WTPT-4','BCUTc-1l','BCUTp-1l','VCH-6','SC-5',
'SPC-6','VP-3','SHsOH','SdO','minHsOH',
'maxHother','maxdO','MAXDP2','ETA_EtaP_F_L','MDEC-23',
'MLFER_A','TopoPSA','WTPT-2','BCUTc-1h','BCUTp-1h',
'SHBd','SHother','SsOH','minHBd','minaaCH',
'minaasC','maxHBd','maxwHBa','maxHBint8','maxHsOH',
'LipoaffinityIndex','ETA_Eta_R_L','MDEO-11','minssCH2','apol',
'ATSc1','ATSm3','SCH-6','VCH-7','maxsOH',
'ETA_dEpsilon_D','ETA_Shape_P','ETA_dBetaP','ATSm2','VC-5',
'SsCH3','SaaO','MLFER_S','WPATH','C1SP2',
'ALogP','ATSc3','ATSc5','minsssN','nBondsD2',
'nsssCH']
Set_feature = Set.loc[:, featuren_all]
print(Set_feature)
X = Set_feature
print(Set_feature)
print(Set_feature.values)
import numpy as np
best_x = [ 1.38253693e+00, -1.27923420e+01 , 3.34890459e-01 , 2.09864822e+00,
-1.37653334e+00, -5.92939907e-01, 2.64158346e+00 ,-1.47876716e+01,
3.46623419e+00 ,1.61801497e+00 ,-2.99576488e+00, -3.48074006e-01,
6.38688156e-01, -5.19686352e-01, 9.00704135e-01, 3.09167458e+00,
-3.47797975e+00 ,-2.37999354e+00, -6.02536699e-01, -3.18153296e+00,
7.42434944e-01,-2.89563644e+00 , 3.20435290e-01 , 2.17453028e+01,
1.03539905e+00, 5.36896266e+00 , 2.09566882e+00, 5.27409742e-01,
3.87202400e+00, 3.36569703e+00 , 6.04737620e+00, 2.61958616e+00,
1.80990651e+00 , 1.43818619e+01 , 1.48127264e+00, 1.60160046e+00,
2.11726944e-01, -2.29679055e+00 ,-3.80125811e+00 , 1.40864553e+00,
1.03629442e+01 ,1.81157543e+01, -2.79026323e+00 ,3.56833458e+00,
2.59651752e+00, 2.47588700e+01 , 1.03023707e+00 , 2.37580593e+00,
-3.24008740e+00, 1.18860580e+00 ,-3.09182989e+00, 1.65976731e+00,
1.67475725e+00 ,2.92308457e+00 ,-4.98709744e-01 , 2.31794159e+00,
7.78201037e+00, 5.95230613e-01, 1.27464854e+00 , 1.11117586e-01,
8.62933321e+00, 7.10097513e+00 ,-1.87948566e+00, 7.08198140e+00,
3.20053469e-02 , 6.00339053e-01 , 1.89878864e+00, 4.70380159e-02,
6.31825754e+00 , 8.39251470e+00, 5.51045242e+00 , 3.68552455e+00,
1.35371005e+01, 3.74846338e+01, 1.66060646e+00, -1.27641030e+01,
3.07137345e+00 ,-2.82324098e+00, 1.55734042e+00, 1.41112778e+01,
8.31509632e+00]
print('------------------------------------------------------------------------------')
print(np.mean(Set2[featuren_all]))
print(np.std(Set2[featuren_all]))
# mix_value.append(i*(np.std(Set2[featuren_all])) + np.mean(Set2[featuren_all]))
best_value = []
for j,i in enumerate(best_x):
best_value.append((i*(np.std(Set2[featuren_all])[j])+(np.mean(Set2[featuren_all])[j])))
print(best_value)
max_list=[]
for i in Set_feature.values:
chazhi = best_x - i
list1=[]
a = 0
for j in chazhi:
if abs(j)<2:
list1.append(abs(j))
elif abs(j)<6:
list1.append(np.sqrt((abs(j))))
elif abs(j)<24:
list1.append((np.sqrt(np.sqrt(abs(j)))))
else:
list1.append(np.sqrt(np.sqrt(np.sqrt(abs(j)))))
a = sum(list1)
max_list.append(a)
arr = np.array(max_list)
mix_index = arr.argsort()[:20][::1].tolist()
mix_index.append(1832)
mix_index.append(1786)
mix_index.append(1784)
mix_index.append(1467)
mix_index.append(1466)
mix_index.append(1464)
mix_index.append(1454)
mix_index.append(515)
mix_index.append(512)
mix_index.append(489)
mix_index.append(478)
mix_index.append(474)
mix_index.append(472)
mix_index.append(470)
mix_index.append(467)
mix_index.append(461)
mix_index.append(460)
Set_d = Set_feature.loc[mix_index].copy()
mix_value = []
for j,i in enumerate(np.min(Set_d)):
mix_value.append(i*(np.std(Set2[featuren_all])[j]) + np.mean(Set2[featuren_all])[j])
max_value = []
for j,i in enumerate(np.max(Set_d)):
max_value.append(i*(np.std(Set2[featuren_all])[j]) + np.mean(Set2[featuren_all])[j])
print(mix_value)
print(max_value)
data = {
'min': mix_value,
'max': max_value,
'fuhao': featuren_all,
}
row_index = featuren_all
col_names=['min', 'max']
df=pd.DataFrame(data,columns=col_names,)
print(df)
df['最优解'] = best_value
df['fuhao'] = featuren_all
print(df)
df.to_csv('E:\\test\jianmo\优化问题问题取值范围DE.csv', index=False)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/setup.py",".py","1033","35","from setuptools import setup, find_packages
from os import path as os_path
import sko
this_directory = os_path.abspath(os_path.dirname(__file__))
# 读取文件内容
def read_file(filename):
with open(os_path.join(this_directory, filename), encoding='utf-8') as f:
long_description = f.read()
return long_description
# 获取依赖
def read_requirements(filename):
return [line.strip() for line in read_file(filename).splitlines()
if not line.startswith('#')]
setup(name='scikit-opt',
python_requires='>=3.5',
version=sko.__version__,
description='Swarm Intelligence in Python',
long_description=read_file('docs/en/README.md'),
long_description_content_type=""text/markdown"",
url='https://github.com/guofei9987/scikit-opt',
author='Guo Fei',
author_email='guofei9987@foxmail.com',
license='MIT',
packages=find_packages(),
platforms=['linux', 'windows', 'macos'],
install_requires=['numpy', 'scipy'],
zip_safe=False)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/CONTRIBUTING.md",".md","2522","62","# Contributing guidelines
This page explains how you can contribute to the development of
scikit-opt by submitting patches, tests, new models, or examples.
scikit-opt is developed on
[Github](https://github.com/guofei9987/scikit-opt) using the
[Git](https://git-scm.com/) version control system.
## Submitting a Bug Report
- Include a short, self-contained code snippet that reproduces the
problem
- Ensure that the bug still exists on latest version.
## Making Changes to the Code
For a pull request to be accepted, you must meet the below requirements.
This greatly helps in keeping the job of maintaining and releasing the
software a shared effort.
- **One branch. One feature.** Branches are cheap and github makes it
easy to merge and delete branches with a few clicks. Avoid the
temptation to lump in a bunch of unrelated changes when working on a
feature, if possible. This helps us keep track of what has changed
when preparing a release.
- Commit messages should be clear and concise. If your commit references or
closes a specific issue, you can close it by mentioning it in the
[commit
message](https://docs.github.com/en/github/managing-your-work-on-github/linking-a-pull-request-to-an-issue).
(*For maintainers*: These suggestions go for Merge commit comments
too. These are partially the record for release notes.)
- Each function, class, method, and attribute needs to be documented.
- If you are adding new functionality, you need to add it to the
documentation by editing (or creating) the appropriate file in
`docs/`.
## How to Submit a Pull Request
So you want to submit a patch to scikit-opt but are not too familiar
with github? Here are the steps you need to take.
1. [Fork](https://help.github.com/articles/fork-a-repo) the
[scikit-opt repository](https://github.com/guofei9987/scikit-opt)
on Github.
2. [Create a new feature
branch](https://git-scm.com/book/en/Git-Branching-Basic-Branching-and-Merging).
Each branch must be self-contained, with a single new feature or
bugfix.
3. Make sure the test suite passes. This includes testing on Python 3.
The easiest way to do this is to either enable
[Travis-CI](https://travis-ci.org/) on your fork, or to make a pull
request and check there.
4. If it is a big, new feature please submit an example.
5. [Submit a pull
request](https://help.github.com/articles/using-pull-requests)
## License
scikit-opt is released under the MIT license.
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/DE.py",".py","3253","100","#!/usr/bin/env python3
# -*- coding: utf-8 -*-
# @Time : 2019/8/20
# @Author : github.com/guofei9987
import numpy as np
from .base import SkoBase
from abc import ABCMeta, abstractmethod
from .operators import crossover, mutation, ranking, selection
from .GA import GeneticAlgorithmBase, GA
class DifferentialEvolutionBase(SkoBase, metaclass=ABCMeta):
pass
class DE(GeneticAlgorithmBase):
def __init__(self, func, n_dim, F=0.5,
size_pop=50, max_iter=200, prob_mut=0.3,
lb=-1, ub=1,
constraint_eq=tuple(), constraint_ueq=tuple()):
super().__init__(func, n_dim, size_pop, max_iter, prob_mut,
constraint_eq=constraint_eq, constraint_ueq=constraint_ueq)
self.F = F
self.V, self.U = None, None
self.lb, self.ub = np.array(lb) * np.ones(self.n_dim), np.array(ub) * np.ones(self.n_dim)
self.crtbp()
def crtbp(self):
# create the population
self.X = np.random.uniform(low=self.lb, high=self.ub, size=(self.size_pop, self.n_dim))
return self.X
def chrom2x(self, Chrom):
pass
def ranking(self):
pass
def mutation(self):
'''
V[i]=X[r1]+F(X[r2]-X[r3]),
where i, r1, r2, r3 are randomly generated
'''
X = self.X
# i is not needed,
# and TODO: r1, r2, r3 should not be equal
random_idx = np.random.randint(0, self.size_pop, size=(self.size_pop, 3))
r1, r2, r3 = random_idx[:, 0], random_idx[:, 1], random_idx[:, 2]
# 这里F用固定值,为了防止早熟,可以换成自适应值
self.V = X[r1, :] + self.F * (X[r2, :] - X[r3, :])
# the lower & upper bound still works in mutation
mask = np.random.uniform(low=self.lb, high=self.ub, size=(self.size_pop, self.n_dim))
self.V = np.where(self.V < self.lb, mask, self.V)
self.V = np.where(self.V > self.ub, mask, self.V)
return self.V
def crossover(self):
'''
if rand < prob_crossover, use V, else use X
'''
mask = np.random.rand(self.size_pop, self.n_dim) < self.prob_mut
self.U = np.where(mask, self.V, self.X)
return self.U
def selection(self):
'''
greedy selection
'''
X = self.X.copy()
f_X = self.x2y().copy()
self.X = U = self.U
f_U = self.x2y()
self.X = np.where((f_X < f_U).reshape(-1, 1), X, U)
return self.X
def run(self, max_iter=None):
self.max_iter = max_iter or self.max_iter
for i in range(self.max_iter):
self.mutation()
self.crossover()
self.selection()
# record the best ones
generation_best_index = self.Y.argmin()
self.generation_best_X.append(self.X[generation_best_index, :].copy())
self.generation_best_Y.append(self.Y[generation_best_index])
self.all_history_Y.append(self.Y)
global_best_index = np.array(self.generation_best_Y).argmin()
global_best_X = self.generation_best_X[global_best_index]
global_best_Y = self.func(np.array([global_best_X]))
return global_best_X, global_best_Y
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/tools.py",".py","1187","49","import numpy as np
def func_transformer(func):
'''
transform this kind of function:
```
def demo_func(x):
x1, x2, x3 = x
return x1 ** 2 + x2 ** 2 + x3 ** 2
```
into this kind of function:
```
def demo_func(x):
x1, x2, x3 = x[:,0], x[:,1], x[:,2]
return x1 ** 2 + (x2 - 0.05) ** 2 + x3 ** 2
```
getting vectorial performance if possible
:param func:
:return:
'''
prefered_function_format = '''
def demo_func(x):
x1, x2, x3 = x[:, 0], x[:, 1], x[:, 2]
return x1 ** 2 + (x2 - 0.05) ** 2 + x3 ** 2
'''
is_vector = getattr(func, 'is_vector', False)
if is_vector:
return func
else:
if func.__code__.co_argcount == 1:
def func_transformed(X):
return np.array([func(x) for x in X])
return func_transformed
elif func.__code__.co_argcount > 1:
def func_transformed(X):
return np.array([func(*tuple(x)) for x in X])
return func_transformed
raise ValueError('''
object function error,
function should be like this:
''' + prefered_function_format)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/__init__.py",".py","345","15","__version__ = '0.5.8'
from . import DE, GA, PSO, SA, ACA, AFSA, IA
def start():
print('''
scikit-opt import successfully,
version: {version}
Author: Guo Fei,
Email: guofei9987@foxmail.com
repo: https://github.com/guofei9987/scikit-opt,
documents: https://scikit-opt.github.io/
'''.format(version=__version__))
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/PSO.py",".py","5725","159","#!/usr/bin/env python3
# -*- coding: utf-8 -*-
# @Time : 2019/8/20
# @Author : github.com/guofei9987
import numpy as np
from scikitopt.sko.tools import func_transformer
from .base import SkoBase
class PSO(SkoBase):
""""""
Do PSO (Particle swarm optimization) algorithm.
This algorithm was adapted from the earlier works of J. Kennedy and
R.C. Eberhart in Particle Swarm Optimization [IJCNN1995]_.
The position update can be defined as:
.. math::
x_{i}(t+1) = x_{i}(t) + v_{i}(t+1)
Where the position at the current step :math:`t` is updated using
the computed velocity at :math:`t+1`. Furthermore, the velocity update
is defined as:
.. math::
v_{ij}(t + 1) = w * v_{ij}(t) + c_{p}r_{1j}(t)[y_{ij}(t) − x_{ij}(t)]
+ c_{g}r_{2j}(t)[\hat{y}_{j}(t) − x_{ij}(t)]
Here, :math:`cp` and :math:`cg` are the cognitive and social parameters
respectively. They control the particle's behavior given two choices: (1) to
follow its *personal best* or (2) follow the swarm's *global best* position.
Overall, this dictates if the swarm is explorative or exploitative in nature.
In addition, a parameter :math:`w` controls the inertia of the swarm's
movement.
.. [IJCNN1995] J. Kennedy and R.C. Eberhart, ""Particle Swarm Optimization,""
Proceedings of the IEEE International Joint Conference on Neural
Networks, 1995, pp. 1942-1948.
Parameters
--------------------
func : function
The func you want to do optimal
dim : int
Number of dimension, which is number of parameters of func.
pop : int
Size of population, which is the number of Particles. We use 'pop' to keep accordance with GA
max_iter : int
Max of iter iterations
Attributes
----------------------
pbest_x : array_like, shape is (pop,dim)
best location of every particle in history
pbest_y : array_like, shape is (pop,1)
best image of every particle in history
gbest_x : array_like, shape is (1,dim)
general best location for all particles in history
gbest_y : float
general best image for all particles in history
gbest_y_hist : list
gbest_y of every iteration
Examples
-----------------------------
see https://scikit-opt.github.io/scikit-opt/#/en/README?id=_3-psoparticle-swarm-optimization
""""""
def __init__(self, func, dim, pop=40, max_iter=150, lb=None, ub=None, w=0.8, c1=0.5, c2=0.5):
self.func = func_transformer(func)
self.w = w # inertia
self.cp, self.cg = c1, c2 # parameters to control personal best, global best respectively
self.pop = pop # number of particles
self.dim = dim # dimension of particles, which is the number of variables of func
self.max_iter = max_iter # max iter
self.has_constraints = not (lb is None and ub is None)
self.lb = -np.ones(self.dim) if lb is None else np.array(lb)
self.ub = np.ones(self.dim) if ub is None else np.array(ub)
assert self.dim == len(self.lb) == len(self.ub), 'dim == len(lb) == len(ub) is not True'
assert np.all(self.ub > self.lb), 'upper-bound must be greater than lower-bound'
self.X = np.random.uniform(low=self.lb, high=self.ub, size=(self.pop, self.dim))
v_high = self.ub - self.lb
self.V = np.random.uniform(low=-v_high, high=v_high, size=(self.pop, self.dim)) # speed of particles
self.Y = self.cal_y() # y = f(x) for all particles
self.pbest_x = self.X.copy() # personal best location of every particle in history
self.pbest_y = self.Y.copy() # best image of every particle in history
self.gbest_x = np.zeros((1, self.dim)) # global best location for all particles
self.gbest_y = np.inf # global best y for all particles
self.gbest_y_hist = [] # gbest_y of every iteration
self.update_gbest()
# record verbose values
self.record_mode = False
self.record_value = {'X': [], 'V': [], 'Y': []}
def update_V(self):
r1 = np.random.rand(self.pop, self.dim)
r2 = np.random.rand(self.pop, self.dim)
self.V = self.w * self.V + \
self.cp * r1 * (self.pbest_x - self.X) + \
self.cg * r2 * (self.gbest_x - self.X)
def update_X(self):
self.X = self.X + self.V
if self.has_constraints:
self.X = np.clip(self.X, self.lb, self.ub)
def cal_y(self):
# calculate y for every x in X
self.Y = self.func(self.X).reshape(-1, 1)
return self.Y
def update_pbest(self):
'''
personal best
:return:
'''
self.pbest_x = np.where(self.pbest_y > self.Y, self.X, self.pbest_x)
self.pbest_y = np.where(self.pbest_y > self.Y, self.Y, self.pbest_y)
def update_gbest(self):
'''
global best
:return:
'''
if self.gbest_y > self.Y.min():
self.gbest_x = self.X[self.Y.argmin(), :].copy()
self.gbest_y = self.Y.min()
def recorder(self):
if not self.record_mode:
return
self.record_value['X'].append(self.X)
self.record_value['V'].append(self.V)
self.record_value['Y'].append(self.Y)
def run(self, max_iter=None):
self.max_iter = max_iter or self.max_iter
for iter_num in range(self.max_iter):
self.update_V()
self.recorder()
self.update_X()
self.cal_y()
self.update_pbest()
self.update_gbest()
self.gbest_y_hist.append(self.gbest_y)
return self
fit = run
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/ACA.py",".py","3436","72","#!/usr/bin/env python3
# -*- coding: utf-8 -*-
# @Time : 2019/9/11
# @Author : github.com/guofei9987
import numpy as np
class ACA_TSP:
def __init__(self, func, n_dim,
size_pop=10, max_iter=20,
distance_matrix=None,
alpha=1, beta=2, rho=0.1,
):
self.func = func
self.n_dim = n_dim # 城市数量
self.size_pop = size_pop # 蚂蚁数量
self.max_iter = max_iter # 迭代次数
self.alpha = alpha # 信息素重要程度
self.beta = beta # 适应度的重要程度
self.rho = rho # 信息素挥发速度
self.prob_matrix_distance = 1 / (distance_matrix + 1e-10 * np.eye(n_dim, n_dim)) # 避免除零错误
self.Tau = np.ones((n_dim, n_dim)) # 信息素矩阵,每次迭代都会更新
self.Table = np.zeros((size_pop, n_dim)).astype(np.int) # 某一代每个蚂蚁的爬行路径
self.y = None # 某一代每个蚂蚁的爬行总距离
self.x_best_history, self.y_best_history = [], [] # 记录各代的最佳情况
self.best_x, self.best_y = None, None
def run(self, max_iter=None):
self.max_iter = max_iter or self.max_iter
for i in range(self.max_iter): # 对每次迭代
prob_matrix = (self.Tau ** self.alpha) * (self.prob_matrix_distance) ** self.beta # 转移概率,无须归一化。
for j in range(self.size_pop): # 对每个蚂蚁
self.Table[j, 0] = 0 # start point,其实可以随机,但没什么区别
for k in range(self.n_dim - 1): # 蚂蚁到达的每个节点
taboo_set = set(self.Table[j, :k + 1]) # 已经经过的点和当前点,不能再次经过
allow_list = list(set(range(self.n_dim)) - taboo_set) # 在这些点中做选择
prob = prob_matrix[self.Table[j, k], allow_list]
prob = prob / prob.sum() # 概率归一化
next_point = np.random.choice(allow_list, size=1, p=prob)[0]
self.Table[j, k + 1] = next_point
# 计算距离
y = np.array([self.func(i) for i in self.Table])
# 顺便记录历史最好情况
index_best = y.argmin()
x_best, y_best = self.Table[index_best, :].copy(), y[index_best].copy()
self.x_best_history.append(x_best)
self.y_best_history.append(y_best)
# 计算需要新涂抹的信息素
delta_tau = np.zeros((self.n_dim, self.n_dim))
for j in range(self.size_pop): # 每个蚂蚁
for k in range(self.n_dim - 1): # 每个节点
n1, n2 = self.Table[j, k], self.Table[j, k + 1] # 蚂蚁从n1节点爬到n2节点
delta_tau[n1, n2] += 1 / y[j] # 涂抹的信息素
n1, n2 = self.Table[j, self.n_dim - 1], self.Table[j, 0] # 蚂蚁从最后一个节点爬回到第一个节点
delta_tau[n1, n2] += 1 / y[j] # 涂抹信息素
# 信息素飘散+信息素涂抹
self.Tau = (1 - self.rho) * self.Tau + delta_tau
best_generation = np.array(self.y_best_history).argmin()
self.best_x = self.x_best_history[best_generation]
self.best_y = self.y_best_history[best_generation]
return self.best_x, self.best_y
fit = run
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/demo_func.py",".py","2888","96","import numpy as np
from scipy import spatial
def function_for_TSP(num_points, seed=None):
if seed:
np.random.seed(seed=seed)
points_coordinate = np.random.rand(num_points, 2) # generate coordinate of points randomly
distance_matrix = spatial.distance.cdist(points_coordinate, points_coordinate, metric='euclidean')
# print('distance_matrix is: \n', distance_matrix)
def cal_total_distance(routine):
num_points, = routine.shape
return sum([distance_matrix[routine[i % num_points], routine[(i + 1) % num_points]] for i in range(num_points)])
return num_points, points_coordinate, distance_matrix, cal_total_distance
def sphere(p):
# Sphere函数
out_put = 0
for i in p:
out_put += i ** 2
return out_put
def schaffer(p):
'''
二维函数,具有无数个极小值点、强烈的震荡形态。很难找到全局最优值
在(0,0)处取的最值0
-10<=x1,x2<=10
'''
x1, x2 = p
x = np.square(x1) + np.square(x2)
return 0.5 + (np.square(np.sin(np.sqrt(x))) - 0.5) / np.square(1 + 0.001 * x)
def shubert(p):
'''
2-dimension
-10<=x1,x2<=10
has 760 local minimas, 18 of which are global minimas with -186.7309
'''
x, y = p
part1 = [i * np.cos((i + 1) * x + i) for i in range(1, 6)]
part2 = [i * np.cos((i + 1) * y + i) for i in range(1, 6)]
return np.sum(part1) * np.sum(part2)
def griewank(p):
'''
存在多个局部最小值点,数目与问题的维度有关。
此函数是典型的非线性多模态函数,具有广泛的搜索空间,是优化算法很难处理的复杂多模态问题。
在(0,...,0)处取的全局最小值0
-600<=xi<=600
'''
part1 = [np.square(x) / 4000 for x in p]
part2 = [np.cos(x / np.sqrt(i + 1)) for i, x in enumerate(p)]
return np.sum(part1) - np.prod(part2) + 1
def rastrigrin(p):
'''
多峰值函数,也是典型的非线性多模态函数
-5.12<=xi<=5.12
在范围内有10n个局部最小值,峰形高低起伏不定跳跃。很难找到全局最优
has a global minimum at x = 0 where f(x) = 0
'''
return np.sum([np.square(x) - 10 * np.cos(2 * np.pi * x) + 10 for x in p])
def rosenbrock(p):
'''
-2.048<=xi<=2.048
函数全局最优点在一个平滑、狭长的抛物线山谷内,使算法很难辨别搜索方向,查找最优也变得十分困难
在(1,...,1)处可以找到极小值0
:param p:
:return:
'''
n_dim = len(p)
res = 0
for i in range(n_dim - 1):
res += 100 * np.square(np.square(p[i]) - p[i + 1]) + np.square(p[i] - 1)
return res
if __name__ == '__main__':
print(sphere((0, 0)))
print(schaffer((0, 0)))
print(shubert((-7.08350643, -7.70831395)))
print(griewank((0, 0, 0)))
print(rastrigrin((0, 0, 0)))
print(rosenbrock((1, 1, 1)))
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/base.py",".py","639","24","from abc import ABCMeta, abstractmethod
import types
class SkoBase(metaclass=ABCMeta):
def register(self, operator_name, operator, *args, **kwargs):
'''
regeister udf to the class
:param operator_name: string
:param operator: a function, operator itself
:param args: arg of operator
:param kwargs: kwargs of operator
:return:
'''
def operator_wapper(*wrapper_args):
return operator(*(wrapper_args + args), **kwargs)
setattr(self, operator_name, types.MethodType(operator_wapper, self))
return self
class Problem(object):
pass","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/AFSA.py",".py","9316","212","#!/usr/bin/env python3
# -*- coding: utf-8 -*-
# @Time : 2019/9/17
# @Author : github.com/guofei9987
import numpy as np
from scipy import spatial
# class ASFA_raw:
# # 这里统一做成极小值求解的问题
# # 参考某 Matlab 教课书上的代码,很多地方的设定无力吐槽,基本没有泛用性
# def __init__(self, func):
# self.func = func
# self.size_pop = 50
# self.max_iter = 300
# self.n_dim = 2
# self.max_try_num = 100 # 最大尝试次数
# self.step = 0.1 # 每一步的最大位移
# self.visual = 0.5 # 鱼的最大感知范围
# self.delta = 1.3 # 拥挤度阈值
#
# self.X = np.random.rand(self.size_pop, self.n_dim)
# self.Y = np.array([self.func(x) for x in self.X])
#
# best_idx = self.Y.argmin()
# self.best_Y = self.Y[best_idx]
# self.best_X = self.X[best_idx, :]
#
# def move_to_target(self, index_individual, x_target):
# # 向目标移动,prey(), swarm(), follow() 三个算子中的移动都用这个
# x = self.X[index_individual, :]
# x_new = x + self.step * np.random.rand() * (x_target - x) / np.linalg.norm(x_target - x)
# self.X[index_individual, :] = x_new
# self.Y[index_individual] = self.func(x_new)
# if self.Y[index_individual] < self.best_Y:
# self.best_X = self.X[index_individual, :]
#
# def move(self, index_individual):
# r = 2 * np.random.rand(self.n_dim) - 1
# x_new = self.X[index_individual, :] + self.visual * r
# self.X[index_individual, :] = x_new
# self.Y[index_individual] = self.func(x_new)
# if self.Y[index_individual] < self.best_Y:
# self.best_X = self.X[index_individual, :]
#
# def prey(self, index_individual):
# for try_num in range(self.max_try_num):
# r = 2 * np.random.rand(self.n_dim) - 1
# x_target = self.X[index_individual, :] + self.visual * r
# if self.func(x_target) < self.Y[index_individual]: # 捕食成功
# print('prey',index_individual)
# self.move_to_target(index_individual, x_target)
# return None
# # 捕食 max_try_num 次后仍不成功,就调用 move 算子
# self.move(index_individual)
#
# def find_individual_in_vision(self, index_individual):
# # 找出 index_individual 这个个体视线范围内的所有鱼
# distances = spatial.distance.cdist(self.X[[index_individual], :], self.X, metric='euclidean').reshape(-1)
# index_individual_in_vision = np.argwhere((distances > 0) & (distances < self.visual))[:, 0]
# return index_individual_in_vision
#
# def swarm(self, index_individual):
# index_individual_in_vision = self.find_individual_in_vision(index_individual)
# num_index_individual_in_vision = len(index_individual_in_vision)
# if num_index_individual_in_vision > 0:
# individual_in_vision = self.X[index_individual_in_vision, :]
# center_individual_in_vision = individual_in_vision.mean(axis=0)
# center_y_in_vision = self.func(center_individual_in_vision)
# if center_y_in_vision * num_index_individual_in_vision < self.delta * self.Y[index_individual]:
# self.move_to_target(index_individual, center_individual_in_vision)
# return None
# self.prey(index_individual)
#
# def follow(self, index_individual):
# index_individual_in_vision = self.find_individual_in_vision(index_individual)
# num_index_individual_in_vision = len(index_individual_in_vision)
# if num_index_individual_in_vision > 0:
# individual_in_vision = self.X[index_individual_in_vision, :]
# y_in_vision = np.array([self.func(x) for x in individual_in_vision])
# index_target = y_in_vision.argmax()
# x_target = individual_in_vision[index_target]
# y_target = y_in_vision[index_target]
# if y_target * num_index_individual_in_vision < self.delta * self.Y[index_individual]:
# self.move_to_target(index_individual, x_target)
# return None
# self.prey(index_individual)
#
# def fit(self):
# for epoch in range(self.max_iter):
# for index_individual in range(self.size_pop):
# self.swarm(index_individual)
# self.follow(index_individual)
# return self.best_X, self.best_Y
# %%
class AFSA:
def __init__(self, func, n_dim, size_pop=50, max_iter=300,
max_try_num=100, step=0.5, visual=0.3,
q=0.98, delta=0.5):
self.func = func
self.n_dim = n_dim
self.size_pop = size_pop
self.max_iter = max_iter
self.max_try_num = max_try_num # 最大尝试捕食次数
self.step = step # 每一步的最大位移比例
self.visual = visual # 鱼的最大感知范围
self.q = q # 鱼的感知范围衰减系数
self.delta = delta # 拥挤度阈值,越大越容易聚群和追尾
self.X = np.random.rand(self.size_pop, self.n_dim)
self.Y = np.array([self.func(x) for x in self.X])
best_idx = self.Y.argmin()
self.best_Y = self.Y[best_idx]
self.best_X = self.X[best_idx, :]
def move_to_target(self, idx_individual, x_target):
'''
move to target
called by prey(), swarm(), follow()
:param idx_individual:
:param x_target:
:return:
'''
x = self.X[idx_individual, :]
x_new = x + self.step * np.random.rand() * (x_target - x)
# x_new = x_target
self.X[idx_individual, :] = x_new
self.Y[idx_individual] = self.func(x_new)
if self.Y[idx_individual] < self.best_Y:
self.best_X = self.X[idx_individual, :].copy()
self.best_Y = self.Y[idx_individual].copy()
def move(self, idx_individual):
'''
randomly move to a point
:param idx_individual:
:return:
'''
r = 2 * np.random.rand(self.n_dim) - 1
x_new = self.X[idx_individual, :] + self.visual * r
self.X[idx_individual, :] = x_new
self.Y[idx_individual] = self.func(x_new)
if self.Y[idx_individual] < self.best_Y:
self.best_X = self.X[idx_individual, :].copy()
self.best_Y = self.Y[idx_individual].copy()
def prey(self, idx_individual):
'''
prey
:param idx_individual:
:return:
'''
for try_num in range(self.max_try_num):
r = 2 * np.random.rand(self.n_dim) - 1
x_target = self.X[idx_individual, :] + self.visual * r
if self.func(x_target) < self.Y[idx_individual]: # 捕食成功
self.move_to_target(idx_individual, x_target)
return None
# 捕食 max_try_num 次后仍不成功,就调用 move 算子
self.move(idx_individual)
def find_individual_in_vision(self, idx_individual):
# 找出 idx_individual 这条鱼视线范围内的所有鱼
distances = spatial.distance.cdist(self.X[[idx_individual], :], self.X, metric='euclidean').reshape(-1)
idx_individual_in_vision = np.argwhere((distances > 0) & (distances < self.visual))[:, 0]
return idx_individual_in_vision
def swarm(self, idx_individual):
# 聚群行为
idx_individual_in_vision = self.find_individual_in_vision(idx_individual)
num_idx_individual_in_vision = len(idx_individual_in_vision)
if num_idx_individual_in_vision > 0:
individual_in_vision = self.X[idx_individual_in_vision, :]
center_individual_in_vision = individual_in_vision.mean(axis=0)
center_y_in_vision = self.func(center_individual_in_vision)
if center_y_in_vision * num_idx_individual_in_vision < self.delta * self.Y[idx_individual]:
self.move_to_target(idx_individual, center_individual_in_vision)
return None
self.prey(idx_individual)
def follow(self, idx_individual):
# 追尾行为
idx_individual_in_vision = self.find_individual_in_vision(idx_individual)
num_idx_individual_in_vision = len(idx_individual_in_vision)
if num_idx_individual_in_vision > 0:
individual_in_vision = self.X[idx_individual_in_vision, :]
y_in_vision = np.array([self.func(x) for x in individual_in_vision])
idx_target = y_in_vision.argmin()
x_target = individual_in_vision[idx_target]
y_target = y_in_vision[idx_target]
if y_target * num_idx_individual_in_vision < self.delta * self.Y[idx_individual]:
self.move_to_target(idx_individual, x_target)
return None
self.prey(idx_individual)
def run(self, max_iter=None):
self.max_iter = max_iter or self.max_iter
for epoch in range(self.max_iter):
for idx_individual in range(self.size_pop):
self.swarm(idx_individual)
self.follow(idx_individual)
self.visual *= self.q
return self.best_X, self.best_Y
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/operators_gpu/mutation_gpu.py",".py","332","14","import torch
def mutation(self):
'''
mutation of 0/1 type chromosome
faster than `self.Chrom = (mask + self.Chrom) % 2`
:param self:
:return:
'''
mask = (torch.rand(size=(self.size_pop, self.len_chrom), device=self.device) < self.prob_mut).type(torch.int8)
self.Chrom ^= mask
return self.Chrom
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/operators_gpu/selection_gpu.py",".py","632","19","import numpy as np
def selection_tournament_faster(self, tourn_size=3):
'''
Select the best individual among *tournsize* randomly chosen
Same with `selection_tournament` but much faster using numpy
individuals,
:param self:
:param tourn_size:
:return:
'''
aspirants_idx = np.random.randint(self.size_pop, size=(self.size_pop, tourn_size))
aspirants_values = self.FitV[aspirants_idx]
winner = aspirants_values.argmax(axis=1) # winner index in every team
sel_index = [aspirants_idx[i, j] for i, j in enumerate(winner)]
self.Chrom = self.Chrom[sel_index, :]
return self.Chrom
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/operators_gpu/__init__.py",".py","0","0","","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/operators_gpu/crossover_gpu.py",".py","605","19","import numpy as np
import torch
def crossover_2point_bit(self):
Chrom, size_pop, len_chrom = self.Chrom, self.size_pop, self.len_chrom
half_size_pop = int(size_pop / 2)
Chrom1, Chrom2 = Chrom[:half_size_pop], Chrom[half_size_pop:]
mask = torch.zeros(size=(half_size_pop, len_chrom), dtype=torch.int8, device=self.device)
for i in range(half_size_pop):
n1, n2 = np.random.randint(0, self.len_chrom, 2)
if n1 > n2:
n1, n2 = n2, n1
mask[i, n1:n2] = 1
mask2 = (Chrom1 ^ Chrom2) & mask
Chrom1 ^= mask2
Chrom2 ^= mask2
return self.Chrom
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/operators_gpu/ranking_gpu.py",".py","0","0","","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/operators/selection.py",".py","2112","66","import numpy as np
def selection_tournament(self, tourn_size=3):
'''
Select the best individual among *tournsize* randomly chosen
individuals,
:param self:
:param tourn_size:
:return:
'''
FitV = self.FitV
sel_index = []
for i in range(self.size_pop):
# aspirants_index = np.random.choice(range(self.size_pop), size=tourn_size)
aspirants_index = np.random.randint(self.size_pop, size=tourn_size)
sel_index.append(max(aspirants_index, key=lambda i: FitV[i]))
self.Chrom = self.Chrom[sel_index, :] # next generation
return self.Chrom
def selection_tournament_faster(self, tourn_size=3):
'''
Select the best individual among *tournsize* randomly chosen
Same with `selection_tournament` but much faster using numpy
individuals,
:param self:
:param tourn_size:
:return:
'''
aspirants_idx = np.random.randint(self.size_pop, size=(self.size_pop, tourn_size))
aspirants_values = self.FitV[aspirants_idx]
winner = aspirants_values.argmax(axis=1) # winner index in every team
sel_index = [aspirants_idx[i, j] for i, j in enumerate(winner)]
self.Chrom = self.Chrom[sel_index, :]
return self.Chrom
def selection_roulette_1(self):
'''
Select the next generation using roulette
:param self:
:return:
'''
FitV = self.FitV
FitV = FitV - FitV.min() + 1e-10
# the worst one should still has a chance to be selected
sel_prob = FitV / FitV.sum()
sel_index = np.random.choice(range(self.size_pop), size=self.size_pop, p=sel_prob)
self.Chrom = self.Chrom[sel_index, :]
return self.Chrom
def selection_roulette_2(self):
'''
Select the next generation using roulette
:param self:
:return:
'''
FitV = self.FitV
FitV = (FitV - FitV.min()) / (FitV.max() - FitV.min() + 1e-10) + 0.2
# the worst one should still has a chance to be selected
sel_prob = FitV / FitV.sum()
sel_index = np.random.choice(range(self.size_pop), size=self.size_pop, p=sel_prob)
self.Chrom = self.Chrom[sel_index, :]
return self.Chrom
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/operators/__init__.py",".py","0","0","","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/operators/ranking.py",".py","429","21","import numpy as np
def ranking(self):
# GA select the biggest one, but we want to minimize func, so we put a negative here
self.FitV = -self.Y
def ranking_linear(self):
'''
For more details see [Baker1985]_.
:param self:
:return:
.. [Baker1985] Baker J E, ""Adaptive selection methods for genetic
algorithms, 1985.
'''
self.FitV = np.argsort(np.argsort(-self.Y))
return self.FitV
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/operators/crossover.py",".py","3328","88","import numpy as np
__all__ = ['crossover_1point', 'crossover_2point', 'crossover_2point_bit', 'crossover_pmx']
def crossover_1point(self):
Chrom, size_pop, len_chrom = self.Chrom, self.size_pop, self.len_chrom
for i in range(0, size_pop, 2):
n = np.random.randint(0, self.len_chrom)
# crossover at the point n
seg1, seg2 = self.Chrom[i, n:].copy(), self.Chrom[i + 1, n:].copy()
self.Chrom[i, n:], self.Chrom[i + 1, n:] = seg2, seg1
return self.Chrom
def crossover_2point(self):
Chrom, size_pop, len_chrom = self.Chrom, self.size_pop, self.len_chrom
for i in range(0, size_pop, 2):
n1, n2 = np.random.randint(0, self.len_chrom, 2)
if n1 > n2:
n1, n2 = n2, n1
# crossover at the points n1 to n2
seg1, seg2 = self.Chrom[i, n1:n2].copy(), self.Chrom[i + 1, n1:n2].copy()
self.Chrom[i, n1:n2], self.Chrom[i + 1, n1:n2] = seg2, seg1
return self.Chrom
def crossover_2point_bit(self):
'''
3 times faster than `crossover_2point`, but only use for 0/1 type of Chrom
:param self:
:return:
'''
Chrom, size_pop, len_chrom = self.Chrom, self.size_pop, self.len_chrom
half_size_pop = int(size_pop / 2)
Chrom1, Chrom2 = Chrom[:half_size_pop], Chrom[half_size_pop:]
mask = np.zeros(shape=(half_size_pop, len_chrom), dtype=int)
for i in range(half_size_pop):
n1, n2 = np.random.randint(0, self.len_chrom, 2)
if n1 > n2:
n1, n2 = n2, n1
mask[i, n1:n2] = 1
mask2 = (Chrom1 ^ Chrom2) & mask
Chrom1 ^= mask2
Chrom2 ^= mask2
return self.Chrom
# def crossover_rv_3(self):
# Chrom, size_pop = self.Chrom, self.size_pop
# i = np.random.randint(1, self.len_chrom) # crossover at the point i
# Chrom1 = np.concatenate([Chrom[::2, :i], Chrom[1::2, i:]], axis=1)
# Chrom2 = np.concatenate([Chrom[1::2, :i], Chrom[0::2, i:]], axis=1)
# self.Chrom = np.concatenate([Chrom1, Chrom2], axis=0)
# return self.Chrom
def crossover_pmx(self):
'''
Executes a partially matched crossover (PMX) on Chrom.
For more details see [Goldberg1985]_.
:param self:
:return:
.. [Goldberg1985] Goldberg and Lingel, ""Alleles, loci, and the traveling
salesman problem"", 1985.
'''
Chrom, size_pop, len_chrom = self.Chrom, self.size_pop, self.len_chrom
for i in range(0, size_pop, 2):
Chrom1, Chrom2 = self.Chrom[i], self.Chrom[i + 1]
cxpoint1, cxpoint2 = np.random.randint(0, self.len_chrom - 1, 2)
if cxpoint1 >= cxpoint2:
cxpoint1, cxpoint2 = cxpoint2, cxpoint1 + 1
# crossover at the point cxpoint1 to cxpoint2
pos1_recorder = {value: idx for idx, value in enumerate(Chrom1)}
pos2_recorder = {value: idx for idx, value in enumerate(Chrom2)}
for j in range(cxpoint1, cxpoint2):
value1, value2 = Chrom1[j], Chrom2[j]
pos1, pos2 = pos1_recorder[value2], pos2_recorder[value1]
Chrom1[j], Chrom1[pos1] = Chrom1[pos1], Chrom1[j]
Chrom2[j], Chrom2[pos2] = Chrom2[pos2], Chrom2[j]
pos1_recorder[value1], pos1_recorder[value2] = pos1, j
pos2_recorder[value1], pos2_recorder[value2] = j, pos2
self.Chrom[i], self.Chrom[i + 1] = Chrom1, Chrom2
return self.Chrom
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/sko/operators/mutation.py",".py","2224","80","import numpy as np
def mutation(self):
'''
mutation of 0/1 type chromosome
faster than `self.Chrom = (mask + self.Chrom) % 2`
:param self:
:return:
'''
#
mask = (np.random.rand(self.size_pop, self.len_chrom) < self.prob_mut)
self.Chrom ^= mask
return self.Chrom
def mutation_TSP_1(self):
'''
every gene in every chromosome mutate
:param self:
:return:
'''
for i in range(self.size_pop):
for j in range(self.n_dim):
if np.random.rand() < self.prob_mut:
n = np.random.randint(0, self.len_chrom, 1)
self.Chrom[i, j], self.Chrom[i, n] = self.Chrom[i, n], self.Chrom[i, j]
return self.Chrom
def swap(individual):
n1, n2 = np.random.randint(0, individual.shape[0] - 1, 2)
if n1 >= n2:
n1, n2 = n2, n1 + 1
individual[n1], individual[n2] = individual[n2], individual[n1]
return individual
def reverse(individual):
'''
Reverse n1 to n2
Also called `2-Opt`: removes two random edges, reconnecting them so they cross
Karan Bhatia, ""Genetic Algorithms and the Traveling Salesman Problem"", 1994
https://pdfs.semanticscholar.org/c5dd/3d8e97202f07f2e337a791c3bf81cd0bbb13.pdf
'''
n1, n2 = np.random.randint(0, individual.shape[0] - 1, 2)
if n1 >= n2:
n1, n2 = n2, n1 + 1
individual[n1:n2] = individual[n1:n2][::-1]
return individual
def transpose(individual):
# randomly generate n1 < n2 < n3. Notice: not equal
n1, n2, n3 = sorted(np.random.randint(0, individual.shape[0] - 2, 3))
n2 += 1
n3 += 2
slice1, slice2, slice3, slice4 = individual[0:n1], individual[n1:n2], individual[n2:n3 + 1], individual[n3 + 1:]
individual = np.concatenate([slice1, slice3, slice2, slice4])
return individual
def mutation_reverse(self):
'''
Reverse
:param self:
:return:
'''
for i in range(self.size_pop):
if np.random.rand() < self.prob_mut:
self.Chrom[i] = reverse(self.Chrom[i])
return self.Chrom
def mutation_swap(self):
for i in range(self.size_pop):
if np.random.rand() < self.prob_mut:
self.Chrom[i] = swap(self.Chrom[i])
return self.Chrom
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/examples/demo_pso.py",".py","537","23","def demo_func(x):
x1, x2, x3 = x
return x1 ** 2 + (x2 - 0.05) ** 2 + x3 ** 2
# %% Do PSO
from sko.PSO import PSO
pso = PSO(func=demo_func, dim=3, pop=40, max_iter=150, lb=[0, -1, 0.5], ub=[1, 1, 1], w=0.8, c1=0.5, c2=0.5)
pso.run()
print('best_x is ', pso.gbest_x, 'best_y is', pso.gbest_y)
# %% Plot the result
import matplotlib.pyplot as plt
plt.plot(pso.gbest_y_hist)
plt.show()
# %% PSO without constraint:
pso = PSO(func=demo_func, dim=3)
fitness = pso.run()
print('best_x is ', pso.gbest_x, 'best_y is', pso.gbest_y)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/examples/demo_de.py",".py","622","34","'''
min f(x1, x2, x3) = x1^2 + x2^2 + x3^2
s.t.
x1*x2 >= 1
x1*x2 <= 5
x2 + x3 = 1
0 <= x1, x2, x3 <= 5
'''
def obj_func(p):
x1, x2, x3 = p
return x1 ** 2 + x2 ** 2 + x3 ** 2
constraint_eq = [
lambda x: 1 - x[1] - x[2]
]
constraint_ueq = [
lambda x: 1 - x[0] * x[1],
lambda x: x[0] * x[1] - 5
]
# %% Do DifferentialEvolution
from sko.DE import DE
de = DE(func=obj_func, n_dim=3, size_pop=50, max_iter=800, lb=[0, 0, 0], ub=[5, 5, 5],
constraint_eq=constraint_eq, constraint_ueq=constraint_ueq)
best_x, best_y = de.run()
print('best_x:', best_x, '\n', 'best_y:', best_y)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/examples/obj_func_demo.py",".py","555","28","import numpy as np
def sphere(p):
# Sphere函数
out_put = 0
for i in p:
out_put += i ** 2
return out_put
def schaffer(p):
'''
这个函数是二维的复杂函数,具有无数个极小值点
在(0,0)处取的最值0
这个函数具有强烈的震荡形态,所以很难找到全局最优质值
:param p:
:return:
'''
x1, x2 = p
x = np.square(x1) + np.square(x2)
return 0.5 + (np.sin(x) - 0.5) / np.square(1 + 0.001 * x)
def myfunc(p):
x = p
return np.sin(x) + np.square(x)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/examples/demo_pso_ani.py",".py","1197","50","# Plot particle history as animation
import numpy as np
from sko.PSO import PSO
def demo_func(x):
x1, x2 = x
return x1 ** 2 + (x2 - 0.05) ** 2
pso = PSO(func=demo_func, dim=2, pop=20, max_iter=40, lb=[-1, -1], ub=[1, 1])
pso.record_mode = True
pso.run()
print('best_x is ', pso.gbest_x, 'best_y is', pso.gbest_y)
# %% Now Plot the animation
import matplotlib.pyplot as plt
from matplotlib.animation import FuncAnimation
record_value = pso.record_value
X_list, V_list = record_value['X'], record_value['V']
fig, ax = plt.subplots(1, 1)
ax.set_title('title', loc='center')
line = ax.plot([], [], 'b.')
X_grid, Y_grid = np.meshgrid(np.linspace(-1.0, 1.0, 40), np.linspace(-1.0, 1.0, 40))
Z_grid = demo_func((X_grid, Y_grid))
ax.contour(X_grid, Y_grid, Z_grid, 20)
ax.set_xlim(-1, 1)
ax.set_ylim(-1, 1)
plt.ion()
p = plt.show()
def update_scatter(frame):
i, j = frame // 10, frame % 10
ax.set_title('iter = ' + str(i))
X_tmp = X_list[i] + V_list[i] * j / 10.0
plt.setp(line, 'xdata', X_tmp[:, 0], 'ydata', X_tmp[:, 1])
return line
ani = FuncAnimation(fig, update_scatter, blit=True, interval=25, frames=300)
plt.show()
# ani.save('pso.gif', writer='pillow')
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/examples/vrp.py",".py","1889","62","import numpy as np
from scipy import spatial
import matplotlib.pyplot as plt
num_customers = 17
num_vehicle = 5
num_points = 1 + num_customers
max_capacity = 5
customers_coordinate = np.random.rand(num_points, 2) # generate coordinate of points
depot_coordinate = np.array([[0.5, 0.5]])
points_coordinate = np.concatenate([depot_coordinate, customers_coordinate], axis=0)
distance_matrix = spatial.distance.cdist(points_coordinate, points_coordinate, metric='euclidean')
def cal_total_distance(routine):
'''The objective function. input routine, return total distance.
cal_total_distance(np.arange(num_points))
'''
num_points, = routine.shape
return distance_matrix[0, routine[0]] \
+ sum([distance_matrix[routine[i % num_points], routine[(i + 1) % num_points]] for i in range(num_points)]) \
+ distance_matrix[routine[-1], 0]
def constraint_capacity(routine):
capacity = 0
c = 0
for i in routine:
if i != 0:
c += 1
else:
capacity = max(capacity, c + 1)
c = 0
capacity = max(capacity, c + 1)
return capacity - max_capacity
# %%
from sko.GA import GA_TSP
ga_tsp = GA_TSP(func=cal_total_distance, n_dim=num_customers, size_pop=50, max_iter=500, prob_mut=1, )
# The index of customers range from 1 to num_customers:
ga_tsp.Chrom = np.concatenate([np.zeros(shape=(ga_tsp.size_pop, num_vehicle - 1), dtype=np.int), ga_tsp.Chrom + 1],
axis=1)
ga_tsp.has_constraint = True
ga_tsp.constraint_ueq = [constraint_capacity]
best_points, best_distance = ga_tsp.run()
# %%
fig, ax = plt.subplots(1, 2)
best_points_ = np.concatenate([[0], best_points, [0]])
best_points_coordinate = points_coordinate[best_points_, :]
ax[0].plot(best_points_coordinate[:, 0], best_points_coordinate[:, 1], 'o-r')
ax[1].plot(ga_tsp.generation_best_Y)
plt.show()
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/_navbar.md",".md","64","4","- Translations
- [:uk: English](/en/)
- [:cn: 中文](/zh/)
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/_coverpage.md",".md","326","16","
# scikit-opt
> Powerful Python module for Heuristic Algorithms
* Genetic Algorithm
* Particle Swarm Optimization
* Simulated Annealing
* Ant Colony Algorithm
* Immune Algorithm
* Artificial Fish Swarm Algorithm
[GitHub](https://github.com/guofei9987/scikit-opt/)
[Get Started](/en/README)
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/_sidebar.md",".md","63","4","
* [English Document](docs/en.md)
* [中文文档](docs/zh.md)
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/make_doc.py",".py","3331","100","# 不想用 Sphinx,也不像弄一堆静态html文件,所以自己写个咯
'''
需要从readme中解析出:
1. ""-> Demo code: [examples/demo_pso.py](examples/demo_pso.py)""
2. 三个```python为开头,三个 ``` 为结尾
3. 从py文件中读出文本,并替换
4. 前几行是求star,只在readme中出现
需要从py文件中解析出:
1. # %% 做断点后赋予index值,然后插入readme
'''
import os
import sys
import re
def search_code(py_file_name, section_idx):
'''
给定py文件名和section序号,返回一个list,内容是py文件中的code(markdown格式)
:param py_file_name:
:param section_idx:
:return:
'''
with open('../' + py_file_name, encoding='utf-8', mode=""r"") as f:
content = f.readlines()
content_new, i, search_idx, idx_first_match = [], 0, 0, None
while i < len(content) and search_idx <= section_idx:
if content[i].startswith('# %%'):
search_idx += 1
i += 1 # 带井号百分号的那一行也跳过去,不要放到文档里面
if search_idx < section_idx:
pass
elif search_idx == section_idx:
idx_first_match = idx_first_match or i # record first match line
content_new.append(content[i])
i += 1
return [
'-> Demo code: [{py_file_name}#s{section_idx}](https://github.com/guofei9987/scikit-opt/blob/master/{py_file_name}#L{idx_first_match})\n'.
format(py_file_name=py_file_name, section_idx=section_idx + 1, idx_first_match=idx_first_match),
'```python\n'] \
+ content_new \
+ ['```\n']
# %%
def make_doc(origin_file):
with open(origin_file, encoding='utf-8', mode=""r"") as f_readme:
readme = f_readme.readlines()
regex = re.compile('\[examples/[\w#.]+\]')
readme_idx = 0
readme_new = []
while readme_idx < len(readme):
readme_line = readme[readme_idx]
if readme_line.startswith('-> Demo code: ['):
# 找到中括号里面的内容,解析为文件名,section号
py_file_name, section_idx = regex.findall(readme[readme_idx])[0][1:-1].split('#s')
section_idx = int(section_idx) - 1
print('插入代码: ', py_file_name, section_idx)
content_new = search_code(py_file_name, section_idx)
readme_new.extend(content_new)
# 往下寻找第一个代码结束位置
while readme[readme_idx] != '```\n':
readme_idx += 1
else:
# 如果不需要插入代码,就用原本的内容
readme_new.append(readme_line)
readme_idx += 1
return readme_new
# 主页 README 和 en/README
readme_new = make_doc(origin_file='../README.md')
with open('../README.md', encoding='utf-8', mode=""w"") as f_readme:
f_readme.writelines(readme_new)
with open('en/README.md', encoding='utf-8', mode=""w"") as f_readme_en:
f_readme_en.writelines(readme_new[20:])
docs = ['zh/README.md',
'zh/more_ga.md', 'en/more_ga.md',
'zh/more_pso.md', 'en/more_pso.md',
'zh/more_sa.md', 'en/more_sa.md',
]
for i in docs:
docs_new = make_doc(origin_file=i)
with open(i, encoding='utf-8', mode=""w"") as f:
f.writelines(docs_new)
sys.exit()
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/en/more_ga.md",".md","3695","82","
## genetic algorithm do integer programming
If you want some variables to be integer, then set the corresponding `precision` to an integer
For example, our objective function is `demo_func`. We want the variables to be integer interval 2, integer interval 1, float. We set `precision=[2, 1, 1e-7]`:
```python
from sko.GA import GA
demo_func = lambda x: (x[0] - 1) ** 2 + (x[1] - 0.05) ** 2 + x[2] ** 2
ga = GA(func=demo_func, n_dim=3, max_iter=500, lb=[-1, -1, -1], ub=[5, 1, 1], precision=[2, 1, 1e-7])
best_x, best_y = ga.run()
print('best_x:', best_x, '\n', 'best_y:', best_y)
```
Notice:
- If `precision` is an integer, the number of all possible value would better be $2^n$, in which case the performance is the best. It also works if the number is not $2^n$
- If `precision` is not an integer, but you still want this mode, manually deal with it. For example, your original `precision=0.5`, just make a new variable, multiplied by `2`
## How to fix start point and end point with GA for TSP
If it is not a cycle graph, no need to do this.
if your start point and end point is (0, 0) and (1, 1). Build up the object function :
- Start point and end point is not the input of the object function. If totally n+2 points including start and end points, the input is the n points.
- And build up the object function, which is the total distance, as actually they are.
```python
import numpy as np
from scipy import spatial
import matplotlib.pyplot as plt
num_points = 20
points_coordinate = np.random.rand(num_points, 2) # generate coordinate of points
start_point=[[0,0]]
end_point=[[1,1]]
points_coordinate=np.concatenate([points_coordinate,start_point,end_point])
distance_matrix = spatial.distance.cdist(points_coordinate, points_coordinate, metric='euclidean')
def cal_total_distance(routine):
'''The objective function. input routine, return total distance.
cal_total_distance(np.arange(num_points))
'''
num_points, = routine.shape
routine = np.concatenate([[num_points], routine, [num_points+1]])
return sum([distance_matrix[routine[i], routine[i + 1]] for i in range(num_points+2-1)])
```
And the same with others:
```python
from sko.GA import GA_TSP
ga_tsp = GA_TSP(func=cal_total_distance, n_dim=num_points, size_pop=50, max_iter=500, prob_mut=1)
best_points, best_distance = ga_tsp.run()
fig, ax = plt.subplots(1, 2)
best_points_ = np.concatenate([[num_points],best_points, [num_points+1]])
best_points_coordinate = points_coordinate[best_points_, :]
ax[0].plot(best_points_coordinate[:, 0], best_points_coordinate[:, 1], 'o-r')
ax[1].plot(ga_tsp.generation_best_Y)
plt.show()
```

For more information, click [here](https://github.com/guofei9987/scikit-opt/issues/58)
## How to set up starting point or initial population
- For `GA`, after `ga=GA(**params)`, use codes like `ga.Chrom = np.random.randint(0,2,size=(80,20))` to manually set the initial population.
- For `DE`, set `de.X` to your initial X.
- For `SA`, there is a parameter `x0`, which is the init point.
- For `PSO`, set `pso.X` to your initial X, and run `pso.cal_y(); pso.update_gbest(); pso.update_pbest()`
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/en/more_sa.md",".md","1926","60","## 3 types of Simulated Annealing
In the ‘fast’ schedule the updates are:
```
u ~ Uniform(0, 1, size = d)
y = sgn(u - 0.5) * T * ((1 + 1/T)**abs(2*u - 1) - 1.0)
xc = y * (upper - lower)
x_new = x_old + xc
c = n * exp(-n * quench)
T_new = T0 * exp(-c * k**quench)
```
In the ‘cauchy’ schedule the updates are:
```
u ~ Uniform(-pi/2, pi/2, size=d)
xc = learn_rate * T * tan(u)
x_new = x_old + xc
T_new = T0 / (1 + k)
```
In the ‘boltzmann’ schedule the updates are:
```
std = minimum(sqrt(T) * ones(d), (upper - lower) / (3*learn_rate))
y ~ Normal(0, std, size = d)
x_new = x_old + learn_rate * y
T_new = T0 / log(1 + k)
```
### Do Simulated Annealing
#### 1. Fast Simulated Annealing
-> Demo code: [examples/demo_sa.py#s4](https://github.com/guofei9987/scikit-opt/blob/master/examples/demo_sa.py#L17)
```python
from sko.SA import SAFast
sa_fast = SAFast(func=demo_func, x0=[1, 1, 1], T_max=1, T_min=1e-9, q=0.99, L=300, max_stay_counter=150)
sa_fast.run()
print('Fast Simulated Annealing: best_x is ', sa_fast.best_x, 'best_y is ', sa_fast.best_y)
```
#### 2. Boltzmann Simulated Annealing
-> Demo code: [examples/demo_sa.py#s5](https://github.com/guofei9987/scikit-opt/blob/master/examples/demo_sa.py#L24)
```python
from sko.SA import SABoltzmann
sa_boltzmann = SABoltzmann(func=demo_func, x0=[1, 1, 1], T_max=1, T_min=1e-9, q=0.99, L=300, max_stay_counter=150)
sa_boltzmann.run()
print('Boltzmann Simulated Annealing: best_x is ', sa_boltzmann.best_x, 'best_y is ', sa_fast.best_y)
```
#### 3. Cauchy Simulated Annealing
-> Demo code: [examples/demo_sa.py#s6](https://github.com/guofei9987/scikit-opt/blob/master/examples/demo_sa.py#L31)
```python
from sko.SA import SACauchy
sa_cauchy = SACauchy(func=demo_func, x0=[1, 1, 1], T_max=1, T_min=1e-9, q=0.99, L=300, max_stay_counter=150)
sa_cauchy.run()
print('Cauchy Simulated Annealing: best_x is ', sa_cauchy.best_x, 'best_y is ', sa_cauchy.best_y)
```
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/en/curve_fitting.md",".md","1153","51","
## curve fitting using GA
Generate toy train datasets
```python
import numpy as np
import matplotlib.pyplot as plt
from sko.GA import GA
x_true = np.linspace(-1.2, 1.2, 30)
y_true = x_true ** 3 - x_true + 0.4 * np.random.rand(30)
plt.plot(x_true, y_true, 'o')
```

Make up residuals
```python
def f_fun(x, a, b, c, d):
return a * x ** 3 + b * x ** 2 + c * x + d
def obj_fun(p):
a, b, c, d = p
residuals = np.square(f_fun(x_true, a, b, c, d) - y_true).sum()
return residuals
```
Do GA
```python
ga = GA(func=obj_fun, n_dim=4, size_pop=100, max_iter=500,
lb=[-2] * 4, ub=[2] * 4)
best_params, residuals = ga.run()
print('best_x:', best_params, '\n', 'best_y:', residuals)
```
Plot the fitting results
```python
y_predict = f_fun(x_true, *best_params)
fig, ax = plt.subplots()
ax.plot(x_true, y_true, 'o')
ax.plot(x_true, y_predict, '-')
plt.show()
```

","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/en/contributors.md",".md","381","11","## contributors
Thanks to those contributors:
- [Bowen Zhang](https://github.com/BalwynZhang)
- [hranYin](https://github.com/hranYin)
- [zhangcogito](https://github.com/zhangcogito)
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/en/speed_up.md",".md","2044","62","## speed up objective function
### Vectorization calculation
If the objective function supports vectorization, it can run much faster.
The following `schaffer1` is an original objective function, `schaffer2` is the corresponding function that supports vectorization operations.
`schaffer2.is_vector = True` is used to tell the algorithm that it supports vectorization operations, otherwise it is non-vectorized by default.
As a result of the operation, the **time cost was reduced to 30%**
```python
import numpy as np
import time
def schaffer1(p):
'''
This function has plenty of local minimum, with strong shocks
global minimum at (0,0) with value 0
'''
x1, x2 = p
x = np.square(x1) + np.square(x2)
return 0.5 + (np.sin(x) - 0.5) / np.square(1 + 0.001 * x)
def schaffer2(p):
'''
This function has plenty of local minimum, with strong shocks
global minimum at (0,0) with value 0
'''
x1, x2 = p[:, 0], p[:, 1]
x = np.square(x1) + np.square(x2)
return 0.5 + (np.sin(x) - 0.5) / np.square(1 + 0.001 * x)
schaffer2.is_vector = True
# %%
from sko.GA import GA
ga1 = GA(func=schaffer1, n_dim=2, size_pop=5000, max_iter=800, lb=[-1, -1], ub=[1, 1], precision=1e-7)
ga2 = GA(func=schaffer2, n_dim=2, size_pop=5000, max_iter=800, lb=[-1, -1], ub=[1, 1], precision=1e-7)
time_start = time.time()
best_x, best_y = ga1.run()
print('best_x:', best_x, '\n', 'best_y:', best_y)
print('time:', time.time() - time_start, ' seconds')
time_start = time.time()
best_x, best_y = ga2.run()
print('best_x:', best_x, '\n', 'best_y:', best_y)
print('time:', time.time() - time_start, ' seconds')
```
output:
>best_x: [-2.98023233e-08 -2.98023233e-08]
best_y: [1.77635684e-15]
time: 88.32313132286072 seconds
>best_x: [2.98023233e-08 2.98023233e-08]
best_y: [1.77635684e-15]
time: 27.68204379081726 seconds
`scikit-opt` still supports non-vectorization, because some functions are difficult to write as vectorization, and some functions are much less readable when vectorized.","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/en/more_pso.md",".md","1637","62","
## demonstrate PSO with animation
step1:do pso
-> Demo code: [examples/demo_pso_ani.py#s1](https://github.com/guofei9987/scikit-opt/blob/master/examples/demo_pso_ani.py#L1)
```python
# Plot particle history as animation
import numpy as np
from sko.PSO import PSO
def demo_func(x):
x1, x2 = x
return x1 ** 2 + (x2 - 0.05) ** 2
pso = PSO(func=demo_func, dim=2, pop=20, max_iter=40, lb=[-1, -1], ub=[1, 1])
pso.record_mode = True
pso.run()
print('best_x is ', pso.gbest_x, 'best_y is', pso.gbest_y)
```
step2: plot animation
-> Demo code: [examples/demo_pso_ani.py#s2](https://github.com/guofei9987/scikit-opt/blob/master/examples/demo_pso_ani.py#L16)
```python
import matplotlib.pyplot as plt
from matplotlib.animation import FuncAnimation
record_value = pso.record_value
X_list, V_list = record_value['X'], record_value['V']
fig, ax = plt.subplots(1, 1)
ax.set_title('title', loc='center')
line = ax.plot([], [], 'b.')
X_grid, Y_grid = np.meshgrid(np.linspace(-1.0, 1.0, 40), np.linspace(-1.0, 1.0, 40))
Z_grid = demo_func((X_grid, Y_grid))
ax.contour(X_grid, Y_grid, Z_grid, 20)
ax.set_xlim(-1, 1)
ax.set_ylim(-1, 1)
plt.ion()
p = plt.show()
def update_scatter(frame):
i, j = frame // 10, frame % 10
ax.set_title('iter = ' + str(i))
X_tmp = X_list[i] + V_list[i] * j / 10.0
plt.setp(line, 'xdata', X_tmp[:, 0], 'ydata', X_tmp[:, 1])
return line
ani = FuncAnimation(fig, update_scatter, blit=True, interval=25, frames=300)
plt.show()
# ani.save('pso.gif', writer='pillow')
```
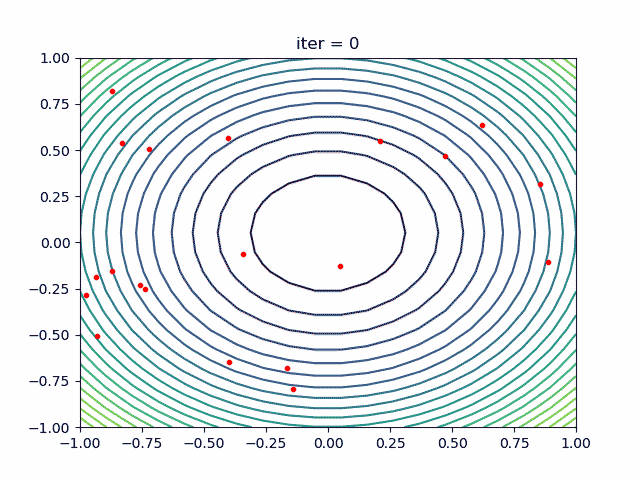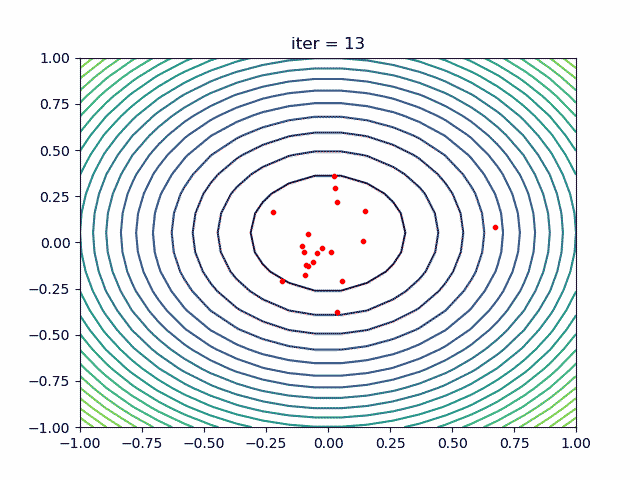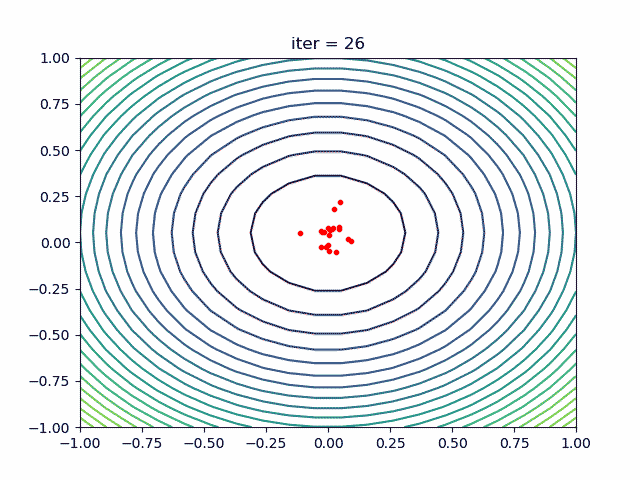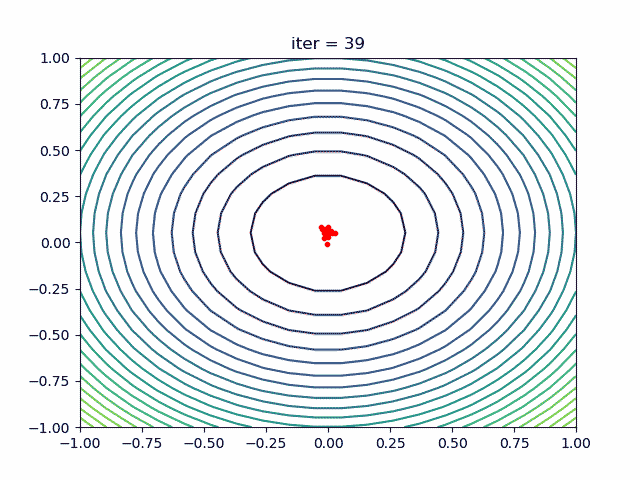 ","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/en/_coverpage.md",".md","351","17","
# scikit-opt
> Powerful Python module for Heuristic Algorithms
* Differential Evolution
* Genetic Algorithm
* Particle Swarm Optimization
* Simulated Annealing
* Ant Colony Algorithm
* Immune Algorithm
* Artificial Fish Swarm Algorithm
[GitHub](https://github.com/guofei9987/scikit-opt/)
[Get Started](/en/README)
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/en/_sidebar.md",".md","241","7","* [English Document](en/README.md)
* [More Genetic Algorithm](en/more_ga.md)
* [More Particle Swarm Optimization](en/more_pso.md)
* [More Simulated Annealing](en/more_sa.md)
* [Curve fiting](en/curve_fitting.md)
* [Speed Up](en/speed_up.md)
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/zh/more_ga.md",".md","3945","83","
## 遗传算法进行整数规划
在多维优化时,想让哪个变量限制为整数,就设定 `precision` 为 **整数** 即可。
例如,我想让我的自定义函数 `demo_func` 的某些变量限制为整数+浮点数(分别是隔2个,隔1个,浮点数),那么就设定 `precision=[2, 1, 1e-7]`
例子如下:
```python
from sko.GA import GA
demo_func = lambda x: (x[0] - 1) ** 2 + (x[1] - 0.05) ** 2 + x[2] ** 2
ga = GA(func=demo_func, n_dim=3, max_iter=500, lb=[-1, -1, -1], ub=[5, 1, 1], precision=[2, 1, 1e-7])
best_x, best_y = ga.run()
print('best_x:', best_x, '\n', 'best_y:', best_y)
```
说明:
- 当 `precision` 为整数时,对应的自变量会启用整数规划模式。
- 在整数规划模式下,变量的取值可能个数最好是 $2^n$,这样收敛速度快,效果好。
- 如果 `precision` 不是整数(例如是0.5),则不会进入整数规划模式,如果还想用这个模式,那么把对应自变量乘以2,这样 `precision` 就是整数了。
## 遗传TSP问题如何固定起点和终点?
固定起点和终点要求路径不闭合(因为如果路径是闭合的,固定与不固定结果实际上是一样的)
假设你的起点和终点坐标指定为(0, 0) 和 (1, 1),这样构建目标函数
- 起点和终点不参与优化。假设共有n+2个点,优化对象是中间n个点
- 目标函数(总距离)按实际去写。
```python
import numpy as np
from scipy import spatial
import matplotlib.pyplot as plt
num_points = 20
points_coordinate = np.random.rand(num_points, 2) # generate coordinate of points
start_point=[[0,0]]
end_point=[[1,1]]
points_coordinate=np.concatenate([points_coordinate,start_point,end_point])
distance_matrix = spatial.distance.cdist(points_coordinate, points_coordinate, metric='euclidean')
def cal_total_distance(routine):
'''The objective function. input routine, return total distance.
cal_total_distance(np.arange(num_points))
'''
num_points, = routine.shape
# start_point,end_point 本身不参与优化。给一个固定的值,参与计算总路径
routine = np.concatenate([[num_points], routine, [num_points+1]])
return sum([distance_matrix[routine[i], routine[i + 1]] for i in range(num_points+2-1)])
```
正常运行并画图:
```python
from sko.GA import GA_TSP
ga_tsp = GA_TSP(func=cal_total_distance, n_dim=num_points, size_pop=50, max_iter=500, prob_mut=1)
best_points, best_distance = ga_tsp.run()
fig, ax = plt.subplots(1, 2)
best_points_ = np.concatenate([[num_points],best_points, [num_points+1]])
best_points_coordinate = points_coordinate[best_points_, :]
ax[0].plot(best_points_coordinate[:, 0], best_points_coordinate[:, 1], 'o-r')
ax[1].plot(ga_tsp.generation_best_Y)
plt.show()
```

更多说明,[这里](https://github.com/guofei9987/scikit-opt/issues/58)
## 如何设定初始点或初始种群
- 对于遗传算法 `GA`, 运行 `ga=GA(**params)` 生成模型后,赋值设定初始种群,例如 `ga.Chrom = np.random.randint(0,2,size=(80,20))`
- 对于差分进化算法 `DE`,设定 `de.X` 为初始 X.
- 对于模拟退火算法 `SA`,入参 `x0` 就是初始点.
- 对于粒子群算法 `PSO`,手动赋值 `pso.X` 为初始 X, 然后执行 `pso.cal_y(); pso.update_gbest(); pso.update_pbest()` 来更新历史最优点
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/zh/more_sa.md",".md","1857","62","## 3 types of Simulated Annealing
模拟退火有三种具体形式
‘fast’:
```
u ~ Uniform(0, 1, size = d)
y = sgn(u - 0.5) * T * ((1 + 1/T)**abs(2*u - 1) - 1.0)
xc = y * (upper - lower)
x_new = x_old + xc
c = n * exp(-n * quench)
T_new = T0 * exp(-c * k**quench)
```
‘cauchy’:
```
u ~ Uniform(-pi/2, pi/2, size=d)
xc = learn_rate * T * tan(u)
x_new = x_old + xc
T_new = T0 / (1 + k)
```
‘boltzmann’:
```
std = minimum(sqrt(T) * ones(d), (upper - lower) / (3*learn_rate))
y ~ Normal(0, std, size = d)
x_new = x_old + learn_rate * y
T_new = T0 / log(1 + k)
```
### 代码示例
#### 1. Fast Simulated Annealing
-> Demo code: [examples/demo_sa.py#s4](https://github.com/guofei9987/scikit-opt/blob/master/examples/demo_sa.py#L17)
```python
from sko.SA import SAFast
sa_fast = SAFast(func=demo_func, x0=[1, 1, 1], T_max=1, T_min=1e-9, q=0.99, L=300, max_stay_counter=150)
sa_fast.run()
print('Fast Simulated Annealing: best_x is ', sa_fast.best_x, 'best_y is ', sa_fast.best_y)
```
#### 2. Boltzmann Simulated Annealing
-> Demo code: [examples/demo_sa.py#s5](https://github.com/guofei9987/scikit-opt/blob/master/examples/demo_sa.py#L24)
```python
from sko.SA import SABoltzmann
sa_boltzmann = SABoltzmann(func=demo_func, x0=[1, 1, 1], T_max=1, T_min=1e-9, q=0.99, L=300, max_stay_counter=150)
sa_boltzmann.run()
print('Boltzmann Simulated Annealing: best_x is ', sa_boltzmann.best_x, 'best_y is ', sa_fast.best_y)
```
#### 3. Cauchy Simulated Annealing
-> Demo code: [examples/demo_sa.py#s6](https://github.com/guofei9987/scikit-opt/blob/master/examples/demo_sa.py#L31)
```python
from sko.SA import SACauchy
sa_cauchy = SACauchy(func=demo_func, x0=[1, 1, 1], T_max=1, T_min=1e-9, q=0.99, L=300, max_stay_counter=150)
sa_cauchy.run()
print('Cauchy Simulated Annealing: best_x is ', sa_cauchy.best_x, 'best_y is ', sa_cauchy.best_y)
```
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/zh/curve_fitting.md",".md","1181","51","
## 使用遗传算法进行曲线拟合
随机生成训练数据
```python
import numpy as np
import matplotlib.pyplot as plt
from sko.GA import GA
x_true = np.linspace(-1.2, 1.2, 30)
y_true = x_true ** 3 - x_true + 0.4 * np.random.rand(30)
plt.plot(x_true, y_true, 'o')
```

构造残差
```python
def f_fun(x, a, b, c, d):
return a * x ** 3 + b * x ** 2 + c * x + d
def obj_fun(p):
a, b, c, d = p
residuals = np.square(f_fun(x_true, a, b, c, d) - y_true).sum()
return residuals
```
使用 scikit-opt 做最优化
```python
ga = GA(func=obj_fun, n_dim=4, size_pop=100, max_iter=500,
lb=[-2] * 4, ub=[2] * 4)
best_params, residuals = ga.run()
print('best_x:', best_params, '\n', 'best_y:', residuals)
```
画出拟合效果图
```python
y_predict = f_fun(x_true, *best_params)
fig, ax = plt.subplots()
ax.plot(x_true, y_true, 'o')
ax.plot(x_true, y_predict, '-')
plt.show()
```

","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/zh/speed_up.md",".md","5417","162","## 目标函数加速
### 矢量化计算
如果目标函数支持矢量化运算,那么运行速度可以大大加快。
下面的 `schaffer1` 是普通的目标函数,`schaffer2` 是支持矢量化运算的目标函数,需要用`schaffer2.is_vector = True`来告诉算法它支持矢量化运算,否则默认是非矢量化的。
从运行结果看,花费时间降低到30%
```python
import numpy as np
import time
def schaffer1(p):
'''
This function has plenty of local minimum, with strong shocks
global minimum at (0,0) with value 0
'''
x1, x2 = p
x = np.square(x1) + np.square(x2)
return 0.5 + (np.sin(x) - 0.5) / np.square(1 + 0.001 * x)
def schaffer2(p):
'''
This function has plenty of local minimum, with strong shocks
global minimum at (0,0) with value 0
'''
x1, x2 = p[:, 0], p[:, 1]
x = np.square(x1) + np.square(x2)
return 0.5 + (np.sin(x) - 0.5) / np.square(1 + 0.001 * x)
schaffer2.is_vector = True
# %%
from sko.GA import GA
ga1 = GA(func=schaffer1, n_dim=2, size_pop=5000, max_iter=800, lb=[-1, -1], ub=[1, 1], precision=1e-7)
ga2 = GA(func=schaffer2, n_dim=2, size_pop=5000, max_iter=800, lb=[-1, -1], ub=[1, 1], precision=1e-7)
time_start = time.time()
best_x, best_y = ga1.run()
print('best_x:', best_x, '\n', 'best_y:', best_y)
print('time:', time.time() - time_start, ' seconds')
time_start = time.time()
best_x, best_y = ga2.run()
print('best_x:', best_x, '\n', 'best_y:', best_y)
print('time:', time.time() - time_start, ' seconds')
```
output:
>best_x: [-2.98023233e-08 -2.98023233e-08]
best_y: [1.77635684e-15]
time: 88.32313132286072 seconds
>best_x: [2.98023233e-08 2.98023233e-08]
best_y: [1.77635684e-15]
time: 27.68204379081726 seconds
`scikit-opt` 仍然支持非矢量化计算,因为有些函数很难写成矢量化计算的形式,还有些函数强行写成矢量化形式后可读性会大大降低。
## 算子优化加速
主要手段是 **矢量化** 和 **逻辑化**
对于可以矢量化的算子,`scikit-opt` 都尽量做了矢量化,并且默认调用矢量化的算子,且 **无须用户额外操作**。
另外,考虑到有些算子矢量化后,代码可读性下降,因此矢量化前的算子也会保留,为用户进阶学习提供方便。
### 0/1 基因的mutation
做一个mask,是一个与 `Chrom` 大小一致的0/1矩阵,如果值为1,那么对应位置进行变异(0变1或1变0)
自然想到用整除2的方式进行
```python
def mutation(self):
# mutation of 0/1 type chromosome
mask = (np.random.rand(self.size_pop, self.len_chrom) < self.prob_mut) * 1
self.Chrom = (mask + self.Chrom) % 2
return self.Chrom
```
如此就实现了一次性对整个种群所有基因变异的矢量化运算。用pycharm的profile功能试了一下,效果良好
再次改进。我还嫌求余数这一步速度慢,画一个真值表
|A|mask:是否变异|A变异后|
|--|--|--|
|1|0|1|
|0|0|0|
|1|1|0|
|0|1|1|
发现这就是一个 **异或**
```python
def mutation2(self):
mask = (np.random.rand(self.size_pop, self.len_chrom) < self.prob_mut)
self.Chrom ^= mask
return self.Chrom
```
测试发现运行速度又快了1~3倍,与最原始的双层循环相比,**快了约20倍**。
### 0/1基因的crossover
同样思路,试试crossover.
- mask同样,1表示对应点交叉,0表示对应点不交叉
做一个真值表,总共8种可能,发现其中只有2种可能基因有变化(等位基因一样时,交叉后的结果与交叉前一样)
|A基因|B基因|是否交叉|交叉后的A基因|交叉后的B基因|
|--|--|--|--|--|
|1|0|1|0|1|
|0|1|1|1|0|
可以用 `异或` 和 `且` 来表示是否变化的表达式: `mask = (A^B)&C`,然后可以计算了`A^=mask, B^=mask`
代码实现
```
def crossover_2point_bit(self):
Chrom, size_pop, len_chrom = self.Chrom, self.size_pop, self.len_chrom
Chrom1, Chrom2 = Chrom[::2], Chrom[1::2]
mask = np.zeros(shape=(int(size_pop / 2),len_chrom),dtype=int)
for i in range(int(size_pop / 2)):
n1, n2 = np.random.randint(0, self.len_chrom, 2)
if n1 > n2:
n1, n2 = n2, n1
mask[i, n1:n2] = 1
mask2 = (Chrom1 ^ Chrom2) & mask
Chrom1^=mask2
Chrom2^=mask2
Chrom[::2], Chrom[1::2]=Chrom1,Chrom2
self.Chrom=Chrom
return self.Chrom
```
测试结果,**效率提升约1倍**。
### 锦标赛选择算子selection_tournament
实战发现,selection_tournament 往往是最耗时的,几乎占用一半时间,因此需要优化。
优化前的算法是遍历,每次选择一组进行锦标赛。但可以在二维array上一次性操作。
```python
def selection_tournament_faster(self, tourn_size=3):
'''
Select the best individual among *tournsize* randomly chosen
Same with `selection_tournament` but much faster using numpy
individuals,
:param self:
:param tourn_size:
:return:
'''
aspirants_idx = np.random.randint(self.size_pop, size=(self.size_pop, tourn_size))
aspirants_values = self.FitV[aspirants_idx]
winner = aspirants_values.argmax(axis=1) # winner index in every team
sel_index = [aspirants_idx[i, j] for i, j in enumerate(winner)]
self.Chrom = self.Chrom[sel_index, :]
return self.Chrom
```
发现own time 和time 都降为原来的10%~15%,**效率提升了约9倍**
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/zh/more_pso.md",".md","1628","62","
## 粒子群算法的动画展示
step1:做pso
-> Demo code: [examples/demo_pso_ani.py#s1](https://github.com/guofei9987/scikit-opt/blob/master/examples/demo_pso_ani.py#L1)
```python
# Plot particle history as animation
import numpy as np
from sko.PSO import PSO
def demo_func(x):
x1, x2 = x
return x1 ** 2 + (x2 - 0.05) ** 2
pso = PSO(func=demo_func, dim=2, pop=20, max_iter=40, lb=[-1, -1], ub=[1, 1])
pso.record_mode = True
pso.run()
print('best_x is ', pso.gbest_x, 'best_y is', pso.gbest_y)
```
step2:画图
-> Demo code: [examples/demo_pso_ani.py#s2](https://github.com/guofei9987/scikit-opt/blob/master/examples/demo_pso_ani.py#L16)
```python
import matplotlib.pyplot as plt
from matplotlib.animation import FuncAnimation
record_value = pso.record_value
X_list, V_list = record_value['X'], record_value['V']
fig, ax = plt.subplots(1, 1)
ax.set_title('title', loc='center')
line = ax.plot([], [], 'b.')
X_grid, Y_grid = np.meshgrid(np.linspace(-1.0, 1.0, 40), np.linspace(-1.0, 1.0, 40))
Z_grid = demo_func((X_grid, Y_grid))
ax.contour(X_grid, Y_grid, Z_grid, 20)
ax.set_xlim(-1, 1)
ax.set_ylim(-1, 1)
plt.ion()
p = plt.show()
def update_scatter(frame):
i, j = frame // 10, frame % 10
ax.set_title('iter = ' + str(i))
X_tmp = X_list[i] + V_list[i] * j / 10.0
plt.setp(line, 'xdata', X_tmp[:, 0], 'ydata', X_tmp[:, 1])
return line
ani = FuncAnimation(fig, update_scatter, blit=True, interval=25, frames=300)
plt.show()
# ani.save('pso.gif', writer='pillow')
```
 ","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/zh/_coverpage.md",".md","298","17","
# scikit-opt
> Powerful Python module for Heuristic Algorithms
* 差分进化算法
* 遗传算法
* 粒子群算法
* 模拟退火算法
* 蚁群算法
* 免疫优化算法
* 鱼群算法
[GitHub](https://github.com/guofei9987/scikit-opt/)
[开始](zh/README)
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题四代码/scikitopt/docs/zh/_sidebar.md",".md","264","8","* [文档](zh/README.md)
* [入参说明](zh/args.md)
* [更多遗传算法](zh/more_ga.md)
* [更多粒子群算法](zh/more_pso.md)
* [更多模拟退火算法](zh/more_sa.md)
* [遗传算法做曲线拟合](zh/curve_fitting.md)
* [提升速度](zh/speed_up.md)
","Markdown"
"ADMET","Bighhhzq/Mathematical-modeling","问题三代码/XGBoost.py",".py","2252","50","import pandas as pd
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from xgboost.sklearn import XGBClassifier
gr = pd.read_csv('./clean451.csv', index_col=0, encoding='gb18030')
Feature_1 = ['ATSm2', 'BCUTc-1h', 'SCH-6', 'VC-5', 'SHBd', 'SsCH3', 'SaaO', 'minHBa', 'hmin', 'LipoaffinityIndex',
'FMF', 'MDEC-23', 'MLFER_S', 'WPATH']
Feature_2 = ['apol', 'ATSc1', 'ATSm3', 'SCH-6', 'VCH-7', 'SP-6', 'SHBd', 'SHsOH', 'SHaaCH', 'minHBa',
'maxsOH', 'ETA_dEpsilon_D', 'ETA_Shape_P', 'ETA_Shape_Y', 'ETA_BetaP_s', 'ETA_dBetaP']
Feature_3 = ['ATSc2', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'SHBd', 'SHother', 'SsOH', 'minHBd', 'minHBa', 'minssCH2',
'minaaCH', 'minaasC', 'maxHBd', 'maxwHBa', 'maxHBint8', 'maxHsOH', 'hmin', 'LipoaffinityIndex',
'ETA_dEpsilon_B', 'ETA_Shape_Y',
'ETA_EtaP_F', 'ETA_Eta_R_L', 'MDEO-11', 'WTPT-4']
Feature_4 = ['ATSc2', 'BCUTc-1l', 'BCUTp-1l', 'VCH-6', 'SC-5', 'SPC-6', 'VP-3', 'SHsOH', 'SdO', 'minHBa',
'minHsOH', 'maxHother', 'maxdO', 'hmin', 'MAXDP2', 'ETA_dEpsilon_B', 'ETA_Shape_Y', 'ETA_EtaP_F_L',
'MDEC-23', 'MLFER_A',
'TopoPSA', 'WTPT-2', 'WTPT-4']
Feature_5 = ['nN', 'ATSc2', 'SCH-7', 'VPC-5', 'SP-6', 'SHaaCH', 'SssCH2', 'SsssCH', 'SssO', 'minHBa',
'mindssC', 'maxsCH3', 'maxsssCH', 'maxssO', 'hmin', 'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y',
'ETA_BetaP', 'ETA_BetaP_s',
'ETA_EtaP_F', 'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'MLFER_E', 'WTPT-4']
feature_df = gr[Feature_2]
x = feature_df.values
# y_var = ['Caco-2', 'CYP3A4', 'hERG', 'hERG', 'MN']
y_var = ['CYP3A4']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=0, train_size=0.7)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
# 拟合XGBoost模型
model = XGBClassifier()
model.fit(x_train, y_train)
# 对测试集做预测
y_pred = model.predict(x_test)
predictions = [round(value) for value in y_pred]
# 评估预测结果
accuracy = accuracy_score(y_test, predictions)
print(""Accuracy: %.2f%%"" % (accuracy * 100.0))
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题三代码/LigheGBM.py",".py","2086","62","import json
import lightgbm as lgb
import pandas as pd
import numpy as np
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
from sklearn import metrics
try:
import cPickle as pickle
except BaseException:
import pickle
gr = pd.read_csv('./clean451.csv', index_col=0, encoding='gb18030')
feature = ['ATSm2', 'ATSm3', 'BCUTc-1h', 'SCH-6', 'VC-5', 'SP-1', 'ECCEN', 'SHBd',
'SsCH3', 'SaaO', 'minHBa', 'minaaO', 'maxaaO', 'hmin',
'LipoaffinityIndex', 'ETA_Beta', 'ETA_Beta_s', 'ETA_Eta_R', 'ETA_Eta_F',
'ETA_Eta_R_L', 'FMF', 'MDEC-12', 'MDEC-23', 'MLFER_S', 'MLFER_E',
'MLFER_L', 'TopoPSA', 'MW', 'WTPT-1', 'WPATH']
feature_df = gr[feature]
x = feature_df.values
print(x)
# y_var = ['Caco-2', 'CYP3A4', 'hERG', 'hERG', 'MN']
y_var = ['Caco-2']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=0, train_size=0.7)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
params = {
'boosting_type': 'gbdt',
'objective': 'multiclass',
'num_class': 7,
'metric': 'multi_error',
'num_leaves': 120,
'min_data_in_leaf': 100,
'learning_rate': 0.06,
'feature_fraction': 0.8,
'bagging_fraction': 0.8,
'bagging_freq': 5,
'lambda_l1': 0.4,
'lambda_l2': 0.5,
'min_gain_to_split': 0.2,
'verbose': -1,
}
print('Training...')
trn_data = lgb.Dataset(x_train, y_train)
val_data = lgb.Dataset(x_test, y_test)
clf = lgb.train(params,
trn_data,
num_boost_round=1000,
valid_sets=[trn_data, val_data],
verbose_eval=100,
early_stopping_rounds=100)
print('Predicting...')
y_prob = clf.predict(x_test, num_iteration=clf.best_iteration)
y_pred = [list(x).index(max(x)) for x in y_prob]
print(""AUC score: {:<8.5f}"".format(metrics.accuracy_score(y_pred, y_test)))
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题三代码/vote.py",".py","3492","87","from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.ensemble import VotingClassifier
from sklearn.pipeline import make_pipeline
from xgboost.sklearn import XGBClassifier
from lightgbm.sklearn import LGBMClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
import pandas as pd
gr = pd.read_csv('./clean451.csv', index_col=0, encoding='gb18030')
Feature_1 = ['ATSm2', 'BCUTc-1h', 'SCH-6', 'VC-5', 'SHBd', 'SsCH3', 'SaaO', 'minHBa', 'hmin', 'LipoaffinityIndex',
'FMF', 'MDEC-23', 'MLFER_S', 'WPATH']
Feature_2 = ['apol', 'ATSc1', 'ATSm3', 'SCH-6', 'VCH-7', 'SP-6', 'SHBd', 'SHsOH', 'SHaaCH', 'minHBa',
'maxsOH', 'ETA_dEpsilon_D', 'ETA_Shape_P', 'ETA_Shape_Y', 'ETA_BetaP_s', 'ETA_dBetaP']
Feature_3 = ['ATSc2', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'SHBd', 'SHother', 'SsOH', 'minHBd', 'minHBa', 'minssCH2',
'minaaCH', 'minaasC', 'maxHBd', 'maxwHBa', 'maxHBint8', 'maxHsOH', 'hmin', 'LipoaffinityIndex',
'ETA_dEpsilon_B', 'ETA_Shape_Y',
'ETA_EtaP_F', 'ETA_Eta_R_L', 'MDEO-11', 'WTPT-4']
Feature_4 = ['ATSc2', 'BCUTc-1l', 'BCUTp-1l', 'VCH-6', 'SC-5', 'SPC-6', 'VP-3', 'SHsOH', 'SdO', 'minHBa',
'minHsOH', 'maxHother', 'maxdO', 'hmin', 'MAXDP2', 'ETA_dEpsilon_B', 'ETA_Shape_Y', 'ETA_EtaP_F_L',
'MDEC-23', 'MLFER_A',
'TopoPSA', 'WTPT-2', 'WTPT-4']
Feature_5 = ['nN', 'ATSc2', 'SCH-7', 'VPC-5', 'SP-6', 'SHaaCH', 'SssCH2', 'SsssCH', 'SssO', 'minHBa',
'mindssC', 'maxsCH3', 'maxsssCH', 'maxssO', 'hmin', 'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y',
'ETA_BetaP', 'ETA_BetaP_s',
'ETA_EtaP_F', 'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'MLFER_E', 'WTPT-4']
feature_df = gr[Feature_5]
x = feature_df.values
# y_var = ['Caco-2', 'CYP3A4', 'hERG', 'HOB', 'MN']
y_var = ['MN']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=0, train_size=0.9)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
rf = RandomForestClassifier()
xgboost = XGBClassifier(eval_metric=['logloss', 'auc', 'error'], use_label_encoder=False)
lgbm = LGBMClassifier()
pipe1 = make_pipeline(StandardScaler(), rf)
pipe2 = make_pipeline(StandardScaler(), xgboost)
pipe3 = make_pipeline(StandardScaler(), lgbm)
models = [
('rf', pipe1),
('xgb', pipe2),
('lgbm', pipe3)
]
ensembel = VotingClassifier(estimators=models, voting='soft')
from sklearn.model_selection import cross_val_score
all_model = [pipe1, pipe2, pipe3, ensembel]
clf_labels = ['RandomForestClassifier', 'XGBClassifier', ""LGBMClassifier"", 'Ensemble']
for clf, label in zip(all_model, clf_labels):
score = cross_val_score(estimator=clf,
X=x_train,
y=y_train,
cv=10,
scoring='roc_auc')
print('roc_auc: %0.4f (+/- %0.2f) [%s]' % (score.mean(), score.std(), label))
clf.fit(x_train, y_train)
print(clf.score(x_test, y_test))
clf = ensembel
## 模型训练
y_pred = clf.fit(x_train, y_train).predict_proba(TT)[:, 1]
pre = y_pred.round()
print(clf.score(x_train, y_train))
print('训练集准确率:', accuracy_score(y_train, clf.predict(x_train)))
print(clf.score(x_test, y_test))
print('测试集准确率:', accuracy_score(y_test, clf.predict(x_test)))
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题三代码/SVM.py",".py","2275","47","import pandas as pd
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
gr = pd.read_csv('./clean451.csv', index_col=0, encoding='gb18030')
Feature_1 = ['ATSm2', 'BCUTc-1h', 'SCH-6', 'VC-5', 'SHBd', 'SsCH3', 'SaaO', 'minHBa', 'hmin', 'LipoaffinityIndex',
'FMF', 'MDEC-23', 'MLFER_S', 'WPATH']
Feature_2 = ['apol', 'ATSc1', 'ATSm3', 'SCH-6', 'VCH-7', 'SP-6', 'SHBd', 'SHsOH', 'SHaaCH', 'minHBa',
'maxsOH', 'ETA_dEpsilon_D', 'ETA_Shape_P', 'ETA_Shape_Y', 'ETA_BetaP_s', 'ETA_dBetaP']
Feature_3 = ['ATSc2', 'BCUTc-1l', 'BCUTc-1h', 'BCUTp-1h', 'SHBd', 'SHother', 'SsOH', 'minHBd', 'minHBa', 'minssCH2',
'minaaCH', 'minaasC', 'maxHBd', 'maxwHBa', 'maxHBint8', 'maxHsOH', 'hmin', 'LipoaffinityIndex',
'ETA_dEpsilon_B', 'ETA_Shape_Y',
'ETA_EtaP_F', 'ETA_Eta_R_L', 'MDEO-11', 'WTPT-4']
Feature_4 = ['ATSc2', 'BCUTc-1l', 'BCUTp-1l', 'VCH-6', 'SC-5', 'SPC-6', 'VP-3', 'SHsOH', 'SdO', 'minHBa',
'minHsOH', 'maxHother', 'maxdO', 'hmin', 'MAXDP2', 'ETA_dEpsilon_B', 'ETA_Shape_Y', 'ETA_EtaP_F_L',
'MDEC-23', 'MLFER_A',
'TopoPSA', 'WTPT-2', 'WTPT-4']
Feature_5 = ['nN', 'ATSc2', 'SCH-7', 'VPC-5', 'SP-6', 'SHaaCH', 'SssCH2', 'SsssCH', 'SssO', 'minHBa',
'mindssC', 'maxsCH3', 'maxsssCH', 'maxssO', 'hmin', 'ETA_dEpsilon_B', 'ETA_dEpsilon_C', 'ETA_Shape_Y',
'ETA_BetaP', 'ETA_BetaP_s',
'ETA_EtaP_F', 'ETA_EtaP_B_RC', 'FMF', 'nHBAcc', 'MLFER_E', 'WTPT-4']
feature_df = gr[Feature_2]
x = feature_df.values
# y_var = ['Caco-2', 'CYP3A4', 'hERG', 'HOB', 'MN']
y_var = ['CYP3A4']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=0, train_size=0.8)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
clf = SVC(C=30, kernel='rbf', decision_function_shape='ovo', max_iter=1000)
## 模型训练
clf.fit(x_train, y_train)
print(clf.score(x_train, y_train))
print('训练集准确率:', accuracy_score(y_train, clf.predict(x_train)))
print(clf.score(x_test, y_test))
print('测试集准确率:', accuracy_score(y_test, clf.predict(x_test)))
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题三代码/Logistic.py",".py","1177","34","from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
import numpy as np
import pandas as pd
gr = pd.read_csv('./data.csv', index_col=0, encoding='gb18030')
# print(gr)
feature = ['ATSm2', 'ATSm3', 'BCUTc-1h', 'SCH-6', 'VC-5', 'SP-1', 'ECCEN', 'SHBd',
'SsCH3', 'SaaO', 'minHBa', 'minaaO', 'maxaaO', 'hmin',
'LipoaffinityIndex', 'ETA_Beta', 'ETA_Beta_s', 'ETA_Eta_R', 'ETA_Eta_F',
'ETA_Eta_R_L', 'FMF', 'MDEC-12', 'MDEC-23', 'MLFER_S', 'MLFER_E',
'MLFER_L', 'TopoPSA', 'MW', 'WTPT-1', 'WPATH']
feature_df = gr[feature]
x = feature_df.values
print(x)
# y_var = ['Caco-2', 'CYP3A4', 'hERG', 'hERG', 'MN']
y_var = ['Caco-2']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=0, train_size=0.7)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
clf = LogisticRegression()
clf = clf.fit(x_train, y_train)
y_predicted = clf.predict(x_test)
accuracy = np.mean(y_predicted == y_test) * 100
print(""y_test\n"", y_test)
print(""y_predicted\n"", y_predicted)
print(""accuracy:"", accuracy)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题三代码/RF.py",".py","1305","38","from sklearn.model_selection import cross_val_score
from sklearn.datasets import load_iris
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
import numpy as np
import pandas as pd
gr = pd.read_csv('./clean451.csv', index_col=0, encoding='gb18030')
# print(gr)
feature = ['ATSm2', 'ATSm3', 'BCUTc-1h', 'SCH-6', 'VC-5', 'SP-1', 'ECCEN', 'SHBd',
'SsCH3', 'SaaO', 'minHBa', 'minaaO', 'maxaaO', 'hmin',
'LipoaffinityIndex', 'ETA_Beta', 'ETA_Beta_s', 'ETA_Eta_R', 'ETA_Eta_F',
'ETA_Eta_R_L', 'FMF', 'MDEC-12', 'MDEC-23', 'MLFER_S', 'MLFER_E',
'MLFER_L', 'TopoPSA', 'MW', 'WTPT-1', 'WPATH']
feature_df = gr[feature]
x = feature_df.values
# x = gr.iloc[:, 6:].values
print(x)
# y_var = ['Caco-2', 'CYP3A4', 'hERG', 'hERG', 'MN']
y_var = ['Caco-2']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=0, train_size=0.7)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
clf = RandomForestClassifier()
clf = clf.fit(x_train, y_train)
y_predicted = clf.predict(x_test)
accuracy = np.mean(y_predicted == y_test) * 100
print(""y_test\n"", y_test)
print(""y_predicted\n"", y_predicted)
print(""accuracy:"", accuracy)
","Python"
"ADMET","Bighhhzq/Mathematical-modeling","问题三代码/BPnetwork.py",".py","3203","110","import pandas as pd
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import classification_report
from sklearn.metrics import recall_score
from sklearn import metrics
import seaborn as sns
from sklearn.model_selection import train_test_split
import torch
import torch.nn.functional as Fun
# defeine BP neural network
class Net1(torch.nn.Module):
def __init__(self, n_feature, n_output=2):
super(Net1, self).__init__()
self.hidden = torch.nn.Linear(n_feature, 50)
self.out = torch.nn.Linear(50, n_output)
def forward(self, x):
x = Fun.relu(self.hidden(x))
x = self.out(x)
return x
class Net2(torch.nn.Module):
def __init__(self, n_feature=729, n_output=2):
super(Net2, self).__init__()
self.hidden1 = torch.nn.Linear(n_feature, 1000)
self.hidden2 = torch.nn.Linear(1000, 200)
self.out = torch.nn.Linear(200, n_output)
def forward(self, x):
x = Fun.relu(self.hidden1(x))
x = Fun.relu(self.hidden2(x))
x = self.out(x)
return x
def printreport(exp, pred):
print(classification_report(exp, pred))
print(""recall score"")
print(recall_score(exp, pred, average='macro'))
gr = pd.read_csv('./clean451.csv', index_col=0, encoding='gb18030')
feature = ['ATSm2', 'ATSm3', 'BCUTc-1h', 'SCH-6', 'VC-5', 'SP-1', 'ECCEN', 'SHBd',
'SsCH3', 'SaaO', 'minHBa', 'minaaO', 'maxaaO', 'hmin',
'LipoaffinityIndex', 'ETA_Beta', 'ETA_Beta_s', 'ETA_Eta_R', 'ETA_Eta_F',
'ETA_Eta_R_L', 'FMF', 'MDEC-12', 'MDEC-23', 'MLFER_S', 'MLFER_E',
'MLFER_L', 'TopoPSA', 'MW', 'WTPT-1', 'WPATH']
feature_df = gr[feature]
x = feature_df.values
print(x)
y_var = ['Caco-2', 'CYP3A4', 'hERG', 'hERG', 'MN']
for v in y_var:
y = gr[v]
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=0, train_size=0.7)
print('训练集和测试集 shape', x_train.shape, y_train.shape, x_test.shape, y_test.shape)
scaler = StandardScaler()
x = scaler.fit_transform(x_train)
input = torch.FloatTensor(x)
label = torch.LongTensor(y_train)
net = Net1(n_feature=30, n_output=2)
optimizer = torch.optim.SGD(net.parameters(), lr=0.05)
# SGD: random gradient decend
loss_func = torch.nn.CrossEntropyLoss()
# define loss function
for i in range(100):
out = net(input)
loss = loss_func(out, label)
optimizer.zero_grad()
# initialize
loss.backward()
optimizer.step()
x = scaler.fit_transform(x_test)
input = torch.FloatTensor(x)
label = torch.Tensor(y_test.to_numpy())
out = net(input)
prediction = torch.max(out, 1)[1]
pred_y = prediction.numpy()
target_y = label.data.numpy()
s = accuracy_score(target_y, pred_y)
print('accury')
print(s)
cm = confusion_matrix(target_y, pred_y)
printreport(target_y, pred_y)
f, ax = plt.subplots(figsize=(5, 5))
sns.heatmap(cm, annot=True, linewidths=0.5, linecolor=""red"", fmt="".0f"", ax=ax)
plt.xlabel(""y_pred"")
plt.ylabel(""y_true"")
plt.show()
","Python"